ramosa2 encodes a LATERAL ORGAN BOUNDARY domain protein that determines the fate of stem cells in branch meristems of maize
- PMID: 16399802
- PMCID: PMC1383634
- DOI: 10.1105/tpc.105.039032
ramosa2 encodes a LATERAL ORGAN BOUNDARY domain protein that determines the fate of stem cells in branch meristems of maize
Abstract
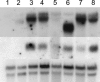
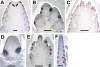
Genetic control of grass inflorescence architecture is critical given that cereal seeds provide most of the world's food. Seeds are borne on axillary branches, which arise from groups of stem cells in axils of leaves and whose branching patterns dictate most of the variation in plant form. Normal maize (Zea mays) ears are unbranched, and tassels have long branches only at their base. The ramosa2 (ra2) mutant of maize has increased branching with short branches replaced by long, indeterminate ones. ra2 was cloned by chromosome walking and shown to encode a LATERAL ORGAN BOUNDARY domain transcription factor. ra2 is transiently expressed in a group of cells that predicts the position of axillary meristem formation in inflorescences. Expression in different mutant backgrounds places ra2 upstream of other genes that regulate branch formation. The early expression of ra2 suggests that it functions in the patterning of stem cells in axillary meristems. Alignment of ra2-like sequences reveals a grass-specific domain in the C terminus that is not found in Arabidopsis thaliana. The ra2-dm allele suggests this domain is required for transcriptional activation of ra1. The ra2 expression pattern is conserved in rice (Oryza sativa), barley (Hordeum vulgare), sorghum (Sorghum bicolor), and maize, suggesting that ra2 is critical for shaping the initial steps of grass inflorescence architecture.
Figures

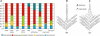


Comment in
-
Branching out: the ramosa pathway and the evolution of grass inflorescence morphology.Plant Cell. 2006 Mar;18(3):518-22. doi: 10.1105/tpc.105.040196. Plant Cell. 2006. PMID: 16513602 Free PMC article. No abstract available.
References
-
- Bonnett, O.T. (1940). Development of the staminate and pistillate inflorescences of sweet corn. J. Agric. Res. 60 25–33.
-
- Booker, J., Sieberer, T., Wright, W., Williamson, L., Willett, B., Stirnberg, P., Turnbull, C., Srinivasan, M., Goddard, P., and Leyser, O. (2005). MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone. Dev. Cell 8 443–449. - PubMed
-
- Chalfun-Junior, A., Franken, J., Mes, J.J., Marsch-Martinez, N., Pereira, A., and Angenent, G.C. (2005). ASYMMETRIC LEAVES2-LIKE1 gene, a member of the AS2/LOB family, controls proximal-distal patterning in Arabidopsis petals. Plant Mol. Biol. 57 559–575. - PubMed
-
- Chuck, G., Muszynski, M., Kellogg, E., Hake, S., and Schmidt, R.J. (2002). The control of spikelet meristem identity by the branched silkless1 gene in maize. Science 298 1238–1241. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

