Identification of novel in vivo Raf-1 phosphorylation sites mediating positive feedback Raf-1 regulation by extracellular signal-regulated kinase
- PMID: 16407412
- PMCID: PMC1382304
- DOI: 10.1091/mbc.e04-12-1123
Identification of novel in vivo Raf-1 phosphorylation sites mediating positive feedback Raf-1 regulation by extracellular signal-regulated kinase
Abstract
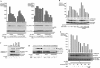
The Ras-Raf-mitogen-activated protein kinase cascade is a key growth-signaling pathway, which uncontrolled activation results in transformation. Although the exact mechanisms underlying Raf-1 regulation remain incompletely understood, phosphorylation has been proposed to play a critical role in this regulation. We report here three novel epidermal growth factor-induced in vivo Raf-1 phosphorylation sites that mediate positive feedback Raf-1 regulation. Using mass spectrometry, we identified Raf-1 phosphorylation on three SP motif sites: S289/S296/S301 and confirmed their identity using two-dimensional-phosphopeptide mapping and phosphospecific antibodies. These sites were phosphorylated by extracellular signal-regulated kinase (ERK)-1 in vitro, and their phosphorylation in vivo was dependent on endogenous ERK activity. Functionally, ERK-1 expression sustains Raf-1 activation in a manner dependent on Raf-1 phosphorylation on the identified sites, and S289/296/301A substitution markedly decreases the in vivo activity of Raf-1 S259A. Importantly, the ERK-phosphorylated Raf-1 pool has 4 times higher specific kinase activity than total Raf-1, and its phosphopeptide composition is similar to that of the general Raf-1 population, suggesting that the preexisting, phosphorylated Raf-1, representing the activatable Raf-1 pool, is the Raf-1 subpopulation targeted by ERK. Our study describes the identification of new in vivo Raf-1 phosphorylation sites targeted by ERK and provides a novel mechanism for a positive feedback Raf-1 regulation.
Figures





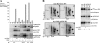
References
-
- Abraham, D., Podar, K., Pacher, M., Kubicek, M., Welzel, N., Hemmings, B. A., Dilworth, S. M., Mischak, H., Kolch, W., and Baccarini, M. (2000). Raf-1-associated protein phosphatase 2A as a positive regulator of kinase activation. J. Biol. Chem. 275, 22300–22304. - PubMed
-
- Alessandrini, A., Greulich, H., Huang, W., and Erikson, R. L. (1996). Mek1 phosphorylation site mutants activate Raf-1 in NIH 3T3 cells. J. Biol. Chem. 271, 31612–31618. - PubMed
-
- Alessi, D. R., Cuenda, A., Cohen, P., Dudley, D. T., and Saltiel, A. R. (1995). PD 098059 is a specific inhibitor of the activation of mitogen-activated protein kinase kinase in vitro and in vivo. J. Biol. Chem. 270, 27489–27494. - PubMed
-
- Avruch, J., Khokhlatchev, A., Kyriakis, J. M., Luo, Z., Tzivion, G., Vavvas, D., and Zhang, X. F. (2001). Ras activation of the Raf kinase: tyrosine kinase recruitment of the MAP kinase cascade. Recent Prog. Horm. Res. 56, 127–155. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

