Two domains that control prefusion stability and transport competence of the measles virus fusion protein
- PMID: 16415028
- PMCID: PMC1346935
- DOI: 10.1128/JVI.80.3.1524-1536.2006
Two domains that control prefusion stability and transport competence of the measles virus fusion protein
Abstract
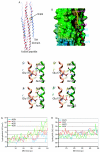
Most viral glycoproteins mediating membrane fusion adopt a metastable native conformation and undergo major conformational changes during fusion. We previously described a panel of compounds that specifically prevent fusion induced by measles virus (MV), most likely by interfering with conformational rearrangements of the MV fusion (F) protein. To further elucidate the basis of inhibition and better understand the mechanism of MV glycoprotein-mediated fusion, we generated and characterized resistant MV variants. Spontaneous mutations conferring drug resistance were confirmed in transient assays and in the context of recombinant virions and were in all cases located in the fusion protein. Several mutations emerged independently at F position 462, which is located in the C-terminal heptad repeat (HR-B) domain. In peptide competition assays, all HR-B mutants at residue 462 revealed reduced affinity for binding to the HR-A core complex compared to unmodified HR-B. Combining mutations at residue 462 with mutations in the distal F head region, which we had previously identified as mediating drug resistance, causes intracellular retention of the mutant proteins. The transport competence and activity of the mutants can be restored, however, by incubation at reduced temperature or in the presence of the inhibitory compounds, indicating that the F escape mutants have a reduced conformational stability and that the inhibitors stabilize a transport-competent conformation of the F trimer. The data support the conclusion that residues located in the head domain of the F trimer and the HR-B region contribute jointly to controlling F conformational stability.
Figures

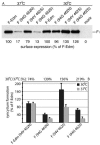

Similar articles
-
Mutations in the putative HR-C region of the measles virus F2 glycoprotein modulate syncytium formation.J Virol. 2003 Apr;77(7):4181-90. doi: 10.1128/jvi.77.7.4181-4190.2003. J Virol. 2003. PMID: 12634376 Free PMC article.
-
Mutations in the Fusion Protein of Measles Virus That Confer Resistance to the Membrane Fusion Inhibitors Carbobenzoxy-d-Phe-l-Phe-Gly and 4-Nitro-2-Phenylacetyl Amino-Benzamide.J Virol. 2017 Nov 14;91(23):e01026-17. doi: 10.1128/JVI.01026-17. Print 2017 Dec 1. J Virol. 2017. PMID: 28904193 Free PMC article.
-
Measles virus entry inhibitors: a structural proposal for mechanism of action and the development of resistance.Biochemistry. 2008 Dec 23;47(51):13573-83. doi: 10.1021/bi801513p. Biochemistry. 2008. PMID: 19035834 Free PMC article.
-
Importance of the cytoplasmic tails of the measles virus glycoproteins for fusogenic activity and the generation of recombinant measles viruses.J Virol. 2002 Jul;76(14):7174-86. doi: 10.1128/jvi.76.14.7174-7186.2002. J Virol. 2002. PMID: 12072517 Free PMC article.
-
A residue located at the junction of the head and stalk regions of measles virus fusion protein regulates membrane fusion by controlling conformational stability.J Gen Virol. 2017 Feb;98(2):143-154. doi: 10.1099/jgv.0.000670. Epub 2017 Mar 13. J Gen Virol. 2017. PMID: 27911256
Cited by
-
Blocking Respiratory Syncytial Virus Entry: A Story with Twists.DNA Cell Biol. 2015 Aug;34(8):505-10. doi: 10.1089/dna.2015.2896. Epub 2015 May 11. DNA Cell Biol. 2015. PMID: 25961744 Free PMC article.
-
Structural Insight into Paramyxovirus and Pneumovirus Entry Inhibition.Viruses. 2020 Mar 20;12(3):342. doi: 10.3390/v12030342. Viruses. 2020. PMID: 32245118 Free PMC article. Review.
-
The F gene of the Osaka-2 strain of measles virus derived from a case of subacute sclerosing panencephalitis is a major determinant of neurovirulence.J Virol. 2010 Nov;84(21):11189-99. doi: 10.1128/JVI.01075-10. Epub 2010 Aug 18. J Virol. 2010. PMID: 20719945 Free PMC article.
-
Residues in the heptad repeat a region of the fusion protein modulate the virulence of Sendai virus in mice.J Virol. 2010 Jan;84(2):810-21. doi: 10.1128/JVI.01990-09. Epub 2009 Nov 11. J Virol. 2010. PMID: 19906935 Free PMC article.
-
A stabilized headless measles virus attachment protein stalk efficiently triggers membrane fusion.J Virol. 2013 Nov;87(21):11693-703. doi: 10.1128/JVI.01945-13. Epub 2013 Aug 21. J Virol. 2013. PMID: 23966411 Free PMC article.
References
-
- Baker, K. A., R. E. Dutch, R. A. Lamb, and T. S. Jardetzky. 1999. Structural basis for paramyxovirus-mediated membrane fusion. Mol. Cell 3:309-319. - PubMed
-
- Ben-Efraim, I., Y. Kliger, C. Hermesh, and Y. Shai. 1999. Membrane-induced step in the activation of Sendai virus fusion protein. J. Mol. Biol. 285:609-625. - PubMed
-
- Berendson, H. J. C., D. Vanderspoel, and R. Vandrunen. 1995. Gromacs—a message-passaging parallel molecular-dynamics implementation. Comp. Phys. Commun. 91:43-56.
-
- Berger, E. A., P. M. Murphy, and J. M. Farber. 1999. Chemokine receptors as HIV-1 coreceptors: roles in viral entry, tropism, and disease. Annu. Rev. Immunol. 17:657-700. - PubMed
-
- Bullough, P. A., F. M. Hughson, J. J. Skehel, and D. C. Wiley. 1994. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371:37-43. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

