Role of protein flexibility in ion permeation: a case study in gramicidin A
- PMID: 16415054
- PMCID: PMC1403166
- DOI: 10.1529/biophysj.105.073205
Role of protein flexibility in ion permeation: a case study in gramicidin A
Abstract
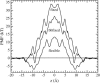
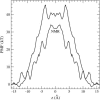
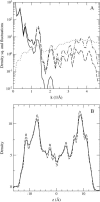
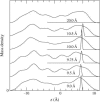
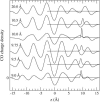
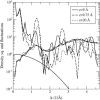
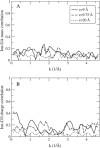
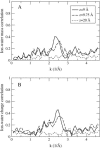
Proteins have a flexible structure, and their atoms exhibit considerable fluctuations under normal operating conditions. However, apart from some enzyme reactions involving ligand binding, our understanding of the role of flexibility in protein function remains mostly incomplete. Here we investigate this question in the realm of membrane proteins that form ion channels. Specifically, we consider ion permeation in the gramicidin A channel, and study how the energetics of ion conduction changes as the channel structure is progressively changed from completely flexible to a fixed one. For each channel structure, the potential of mean force for a permeating potassium ion is determined from molecular dynamics (MD) simulations. Using the same molecular dynamics data for completely flexible gramicidin A, we also calculate the average densities and fluctuations of the peptide atoms and investigate the correlations between these fluctuations and the motion of a permeating ion. Our results show conclusively that peptide flexibility plays an important role in ion permeation in the gramicidin A channel, thus providing another reason--besides the well-known problem with the description of single file pore water--why this channel cannot be modeled using continuum electrostatics with a fixed structure. The new method developed here for studying the role of protein flexibility on its function clarifies the contributions of the fluctuations to energy and entropy, and places limits on the level of detail required in a coarse-grained model.
Figures



Similar articles
-
Test of molecular dynamics force fields in gramicidin A.Eur Biophys J. 2005 Jul;34(5):377-82. doi: 10.1007/s00249-005-0463-2. Epub 2005 Feb 12. Eur Biophys J. 2005. PMID: 15711809
-
Continuum electrostatics fails to describe ion permeation in the gramicidin channel.Biophys J. 2002 Sep;83(3):1348-60. doi: 10.1016/S0006-3495(02)73905-2. Biophys J. 2002. PMID: 12202360 Free PMC article.
-
Gramicidin A channel as a test ground for molecular dynamics force fields.Biophys J. 2003 Apr;84(4):2159-68. doi: 10.1016/S0006-3495(03)75022-X. Biophys J. 2003. PMID: 12668425 Free PMC article.
-
Theoretical and computational models of biological ion channels.Q Rev Biophys. 2004 Feb;37(1):15-103. doi: 10.1017/s0033583504003968. Q Rev Biophys. 2004. PMID: 17390604 Review.
-
Gramicidin channels.IEEE Trans Nanobioscience. 2005 Mar;4(1):10-20. doi: 10.1109/tnb.2004.842470. IEEE Trans Nanobioscience. 2005. PMID: 15816168 Review.
Cited by
-
Computational Investigation of the Effect of Lipid Membranes on Ion Permeation in Gramicidin A.Membranes (Basel). 2016 Mar 18;6(1):20. doi: 10.3390/membranes6010020. Membranes (Basel). 2016. PMID: 26999229 Free PMC article.
-
Site-specific ion occupation in the selectivity filter causes voltage-dependent gating in a viral K+ channel.Sci Rep. 2018 Jul 10;8(1):10406. doi: 10.1038/s41598-018-28751-w. Sci Rep. 2018. PMID: 29991721 Free PMC article.
-
Design of peptide-membrane interactions to modulate single-file water transport through modified gramicidin channels.Biophys J. 2012 Oct 17;103(8):1698-705. doi: 10.1016/j.bpj.2012.08.059. Epub 2012 Oct 16. Biophys J. 2012. PMID: 23083713 Free PMC article.
-
Gaining ligand selectivity in thyroid hormone receptors via entropy.Proc Natl Acad Sci U S A. 2009 Dec 8;106(49):20717-22. doi: 10.1073/pnas.0911024106. Epub 2009 Nov 19. Proc Natl Acad Sci U S A. 2009. PMID: 19926848 Free PMC article.
-
Conduction properties of KcsA measured using brownian dynamics with flexible carbonyl groups in the selectivity filter.Biophys J. 2007 Jul 1;93(1):44-53. doi: 10.1529/biophysj.106.098954. Epub 2007 Apr 13. Biophys J. 2007. PMID: 17434934 Free PMC article.
References
-
- Fersht, A. 1999. Structure and Mechanism in Protein Science. Freeman, New York.
-
- Austin, R. H., K. W. Beeson, L. Eisenstein, and H. Frauenfelder. 1975. Dynamics of ligand binding to myoglobin. Biochemistry. 14:5355–5373. - PubMed
-
- McCammon, J. A., B. R. Gelin, and M. Karplus. 1977. Dynamics of folded proteins. Nature. 267:585–590. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

