Mechanism of polymerase II transcription repression by the histone variant macroH2A
- PMID: 16428466
- PMCID: PMC1347033
- DOI: 10.1128/MCB.26.3.1156-1164.2006
Mechanism of polymerase II transcription repression by the histone variant macroH2A
Abstract
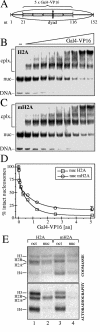
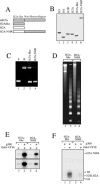
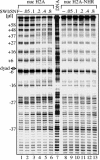
macroH2A (mH2A) is an unusual histone variant consisting of a histone H2A-like domain fused to a large nonhistone region. In this work, we show that histone mH2A represses p300- and Gal4-VP16-dependent polymerase II transcription, and we have dissected the mechanism by which this repression is realized. The repressive effect of mH2A is observed at the level of initiation but not at elongation of transcription, and mH2A interferes with p300-dependent histone acetylation. The nonhistone region of mH2A is responsible for both the repression of initiation of transcription and the inhibition of histone acetylation. In addition, the presence of this domain of mH2A within the nucleosome is able to block nucleosome remodeling and sliding of the histone octamer to neighboring DNA segments by the remodelers SWI/SNF and ACF. These data unambiguously identify mH2A as a strong transcriptional repressor and show that the repressive effect of mH2A is realized on at least two different transcription activation chromatin-dependent pathways: histone acetylation and nucleosome remodeling.
Figures




References
-
- Allen, M. D., A. M. Buckle, S. C. Cordell, J. Lowe, and M. Bycroft. 2003. The crystal structure of AF1521 a protein from Archaeoglobus fulgidus with homology to the non-histone domain of macroH2A. J. Mol. Biol. 330:503-511. - PubMed
-
- An, W., V. B. Palhan, M. A. Karymov, S. H. Leuba, and R. G. Roeder. 2002. Selective requirements for histone H3 and H4 N termini in p300-dependent transcriptional activation from chromatin. Mol. Cell 9:811-821. - PubMed
-
- An, W., and R. G. Roeder. 2003. Direct association of p300 with unmodified H3 and H4 N termini modulates p300-dependent acetylation and transcription of nucleosomal templates. J. Biol. Chem. 278:1504-1510. - PubMed
-
- Angelov, D., A. Molla, P. Y. Perche, F. Hans, J. Cote, S. Khochbin, P. Bouvet, and S. Dimitrov. 2003. The histone variant macroH2A interferes with transcription factor binding and SWI/SNF nucleosome remodeling. Mol. Cell 11:1033-1041. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
