Genetic analysis of completely sequenced disease-associated MHC haplotypes identifies shuffling of segments in recent human history
- PMID: 16440057
- PMCID: PMC1331980
- DOI: 10.1371/journal.pgen.0020009
Genetic analysis of completely sequenced disease-associated MHC haplotypes identifies shuffling of segments in recent human history
Abstract
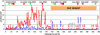
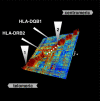
The major histocompatibility complex (MHC) is recognised as one of the most important genetic regions in relation to common human disease. Advancement in identification of MHC genes that confer susceptibility to disease requires greater knowledge of sequence variation across the complex. Highly duplicated and polymorphic regions of the human genome such as the MHC are, however, somewhat refractory to some whole-genome analysis methods. To address this issue, we are employing a bacterial artificial chromosome (BAC) cloning strategy to sequence entire MHC haplotypes from consanguineous cell lines as part of the MHC Haplotype Project. Here we present 4.25 Mb of the human haplotype QBL (HLA-A26-B18-Cw5-DR3-DQ2) and compare it with the MHC reference haplotype and with a second haplotype, COX (HLA-A1-B8-Cw7-DR3-DQ2), that shares the same HLA-DRB1, -DQA1, and -DQB1 alleles. We have defined the complete gene, splice variant, and sequence variation contents of all three haplotypes, comprising over 259 annotated loci and over 20,000 single nucleotide polymorphisms (SNPs). Certain coding sequences vary significantly between different haplotypes, making them candidates for functional and disease-association studies. Analysis of the two DR3 haplotypes allowed delineation of the shared sequence between two HLA class II-related haplotypes differing in disease associations and the identification of at least one of the sites that mediated the original recombination event. The levels of variation across the MHC were similar to those seen for other HLA-disparate haplotypes, except for a 158-kb segment that contained the HLA-DRB1, -DQA1, and -DQB1 genes and showed very limited polymorphism compatible with identity-by-descent and relatively recent common ancestry (<3,400 generations). These results indicate that the differential disease associations of these two DR3 haplotypes are due to sequence variation outside this central 158-kb segment, and that shuffling of ancestral blocks via recombination is a potential mechanism whereby certain DR-DQ allelic combinations, which presumably have favoured immunological functions, can spread across haplotypes and populations.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures



References
-
- Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature. 2001;409:928–933. - PubMed
-
- The International HapMap Project. The International HapMap Project. Nature. 2003;426:789–796. - PubMed
-
- Allcock RJ, Atrazhev AM, Beck S, de Jong PJ, Elliott JF, et al. The MHC haplotype project: A resource for HLA-linked association studies. Tissue Antigens. 2002;59:520–521. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

