Target hub proteins serve as master regulators of development in yeast
- PMID: 16449570
- PMCID: PMC1369046
- DOI: 10.1101/gad.1389306
Target hub proteins serve as master regulators of development in yeast
Abstract
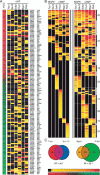
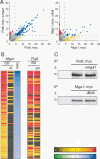
To understand the organization of the transcriptional networks that govern cell differentiation, we have investigated the transcriptional circuitry controlling pseudohyphal development in Saccharomyces cerevisiae. The binding targets of Ste12, Tec1, Sok2, Phd1, Mga1, and Flo8 were globally mapped across the yeast genome. The factors and their targets form a complex binding network, containing patterns characteristic of autoregulation, feedback and feed-forward loops, and cross-talk. Combinatorial binding to intergenic regions was commonly observed, which allowed for the identification of a novel binding association between Mga1 and Flo8, in which Mga1 requires Flo8 for binding to promoter regions. Further analysis of the network showed that the promoters of MGA1 and PHD1 were bound by all of the factors used in this study, identifying them as key target hubs. Overexpression of either of these two proteins specifically induced pseudohyphal growth under noninducing conditions, highlighting them as master regulators of the system. Our results indicate that target hubs can serve as master regulators whose activity is sufficient for the induction of complex developmental responses and therefore represent important regulatory nodes in biological networks.
Figures




References
-
- Barabasi A.L. and Oltvai, Z.N. 2004. Network biology: Understanding the cell's functional organization. Nat. Rev. Genet. 5: 101–113. - PubMed
-
- Boyle E.I., Weng, S., Gollub, J., Jin, H., Botstein, D., Cherry, J.M., and Sherlock, G. 2004. GO::TermFinder: Open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 20: 3710–3715. - PMC - PubMed
-
- Chae J.H., Stein, G.H., and Lee, J.E. 2004. NeuroD: The predicted and the surprising. Mol. Cell 18: 271–288. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
