Tetrapyrrole biosynthesis in Rhodobacter capsulatus is transcriptionally regulated by the heme-binding regulatory protein, HbrL
- PMID: 16452440
- PMCID: PMC1367214
- DOI: 10.1128/JB.188.4.1567-1576.2006
Tetrapyrrole biosynthesis in Rhodobacter capsulatus is transcriptionally regulated by the heme-binding regulatory protein, HbrL
Abstract
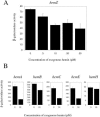
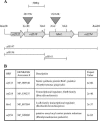
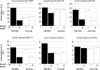
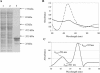
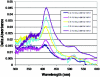
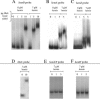
We demonstrate that the expression of hem genes in Rhodobacter capsulatus is transcriptionally repressed in response to the exogenous addition of heme. A high-copy suppressor screen for regulators of hem gene expression resulted in the identification of an LysR-type transcriptional regulator, called HbrL, that regulates hem promoters in response to the availability of heme. HbrL is shown to activate the expression of hemA and hemZ in the absence of exogenous hemin and repress hemB expression in the presence of exogenous hemin. Heterologously expressed HbrL apoprotein binds heme b and is purified with bound heme b when expressed in the presence of 5-aminolevulinic acid. Electrophoretic gel shift analysis demonstrated that HbrL binds the promoter region of hemA, hemB, and hemZ as well as its own promoter and that the presence of heme increases the binding affinity of HbrL to hemB.
Figures

Similar articles
-
The LysR-type transcription factor HbrL is a global regulator of iron homeostasis and porphyrin synthesis in Rhodobacter capsulatus.Mol Microbiol. 2013 Dec;90(6):1277-92. doi: 10.1111/mmi.12431. Epub 2013 Nov 8. Mol Microbiol. 2013. PMID: 24134691 Free PMC article.
-
Regulation of hem gene expression in Rhodobacter capsulatus by redox and photosystem regulators RegA, CrtJ, FnrL, and AerR.J Mol Biol. 2004 Sep 24;342(4):1171-86. doi: 10.1016/j.jmb.2004.08.007. J Mol Biol. 2004. PMID: 15351643
-
The tetrapyrrole biosynthetic pathway and its regulation in Rhodobacter capsulatus.Adv Exp Med Biol. 2010;675:229-50. doi: 10.1007/978-1-4419-1528-3_13. Adv Exp Med Biol. 2010. PMID: 20532744 Free PMC article. Review.
-
The Rhodobacter capsulatus glnB gene is regulated by NtrC at tandem rpoN-independent promoters.J Bacteriol. 1994 Aug;176(16):5171-6. doi: 10.1128/jb.176.16.5171-5176.1994. J Bacteriol. 1994. PMID: 8051036 Free PMC article.
-
Control of photosystem genes in Rhodobacter capsulatus.Trends Genet. 1993 Feb;9(2):56-60. doi: 10.1016/0168-9525(93)90188-N. Trends Genet. 1993. PMID: 8456503 Review.
Cited by
-
Controlling the delicate balance of tetrapyrrole biosynthesis.Philos Trans R Soc Lond B Biol Sci. 2013 Jun 10;368(1622):20120262. doi: 10.1098/rstb.2012.0262. Print 2013 Jul 19. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 23754814 Free PMC article. Review.
-
The LysR-type transcription factor HbrL is a global regulator of iron homeostasis and porphyrin synthesis in Rhodobacter capsulatus.Mol Microbiol. 2013 Dec;90(6):1277-92. doi: 10.1111/mmi.12431. Epub 2013 Nov 8. Mol Microbiol. 2013. PMID: 24134691 Free PMC article.
-
Prokaryotic Heme Biosynthesis: Multiple Pathways to a Common Essential Product.Microbiol Mol Biol Rev. 2017 Jan 25;81(1):e00048-16. doi: 10.1128/MMBR.00048-16. Print 2017 Mar. Microbiol Mol Biol Rev. 2017. PMID: 28123057 Free PMC article. Review.
-
The ChrA-ChrS and HrrA-HrrS signal transduction systems are required for activation of the hmuO promoter and repression of the hemA promoter in Corynebacterium diphtheriae.Infect Immun. 2007 May;75(5):2421-31. doi: 10.1128/IAI.01821-06. Epub 2007 Mar 12. Infect Immun. 2007. PMID: 17353293 Free PMC article.
-
Heme homeostasis and its regulation by hemoproteins in bacteria.mLife. 2024 Jul 11;3(3):327-342. doi: 10.1002/mlf2.12120. eCollection 2024 Sep. mLife. 2024. PMID: 39359680 Free PMC article. Review.
References
-
- Akakura, R., and S. C. Winans. 2002. Constitutive mutations of the OccR regulatory protein affect DNA bending in response to metabolites released from plant tumors. J. Biol. Chem. 277:5866-5874. - PubMed
-
- Akakura, R., and S. C. Winans. 2002. Mutations in the occQ operator that decrease OccR-induced DNA bending do not cause constitutive promoter activity. J. Biol. Chem. 277:15773-15780. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Barany, F. 1985. Single-stranded hexameric linkers: a system for in-phase insertion mutagenesis and protein engineering. Gene 37:111-123. - PubMed
-
- Bender, R. A. 1991. The role of the NAC protein in the nitrogen regulation of Klebsiella aerogenes. Mol. Microbiol. 5:2575-2580. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

