Rhodamine-RCA in vivo labeling guided laser capture microdissection of cancer functional angiogenic vessels in a murine squamous cell carcinoma mouse model
- PMID: 16457726
- PMCID: PMC1420324
- DOI: 10.1186/1476-4598-5-5
Rhodamine-RCA in vivo labeling guided laser capture microdissection of cancer functional angiogenic vessels in a murine squamous cell carcinoma mouse model
Abstract
Background: Cancer growth, invasion and metastasis are highly related to tumor-associated neovasculature. The presence and progression of endothelial cells in cancer is chaotic, unorganized, and angiogenic vessels are less functional. Therefore, not all markers appearing on the chaotic endothelial cells are accessible if a drug is given through the vascular route. Identifying endothelial cell markers from functional cancer angiogenic vessels will indicate the accessibility and potential efficacy of vascular targeted therapies.
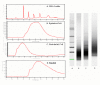
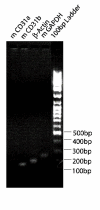
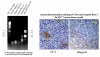
Results: In order to quickly and effectively identify endothelial cell markers on the functional and accessible tumor vessels, we developed a novel technique by which tumor angiogenic vessels are labeled in vivo followed by Laser Capture Microdissection of microscopically isolated endothelial cells for genomic screening. Female C3H mice (N = 5) with established SCCVII tumors were treated with Rhodamine-RCA lectin by tail vein injection, and after fluorescence microscopy showed a successful vasculature staining, LCM was then performed on frozen section tissue using the PixCell II instrument with CapSure HS caps under the Rhodamine filter. By this approach, the fluorescent angiogenic endothelial cells were successfully picked up. As a result, the total RNA concentration increased from an average of 33.4 ng/ul +/- 24.3 (mean +/- S.D.) to 1913.4 ng/ul +/- 164. Relatively pure RNA was retrieved from both endothelial and epithelial cells as indicated by the 260/280 ratios (range 2.22-2.47). RT-PCR and gene electrophoresis successfully detected CD31 and Beta-Actin molecules with minimal Keratin 19 expression, which served as the negative control.
Conclusion: Our present study demonstrates that in vivo Rhodamine RCA angiogenic vessel labeling provided a practical approach to effectively guide functional endothelial cell isolation by laser capture microdissection with fluorescent microscopy, resulting in high quality RNA and pure samples of endothelial cells pooled for detecting genomic expression.
Figures



Similar articles
-
Assessment of gene expression in head and neck carcinoma using laser capture microdissection and real-time reverse transcription polymerase chain reaction.Laryngoscope. 2004 Dec;114(12):2123-8. doi: 10.1097/01.mlg.0000149446.14770.52. Laryngoscope. 2004. PMID: 15564832
-
Development of rapid staining protocols for laser-capture microdissection of brain vessels from human and rat coupled to gene expression analyses.J Neurosci Methods. 2004 Feb 15;133(1-2):39-48. doi: 10.1016/j.jneumeth.2003.09.026. J Neurosci Methods. 2004. PMID: 14757343
-
Human esophageal cancer endothelial cells increase tumor growth by incorporating with mouse endothelium.Cancer Lett. 2007 Jul 8;252(1):123-30. doi: 10.1016/j.canlet.2006.12.015. Epub 2007 Feb 5. Cancer Lett. 2007. PMID: 17276590
-
Application of laser-capture microdissection to analysis of gene expression in the testis.Prog Histochem Cytochem. 2008;42(4):173-201. doi: 10.1016/j.proghi.2007.10.001. Prog Histochem Cytochem. 2008. PMID: 18243898 Review.
-
Tumor vascular biomarkers: new opportunities for cancer diagnostics.Cancer Biomark. 2010-2011;8(4-5):253-71. doi: 10.3233/CBM-2011-0217. Cancer Biomark. 2010. PMID: 22045357 Review.
Cited by
-
Differential Lectin Binding Patterns Identify Distinct Heart Regions in Giant Danio ( Devario aequipinnatus) and Zebrafish ( Danio rerio) Hearts.J Histochem Cytochem. 2016 Nov;64(11):687-714. doi: 10.1369/0022155416667928. Epub 2016 Sep 30. J Histochem Cytochem. 2016. PMID: 27680670 Free PMC article.
-
Tumor angiogenic endothelial cell targeting by a novel integrin-targeted nanoparticle.Int J Nanomedicine. 2007;2(3):479-85. Int J Nanomedicine. 2007. PMID: 18019845 Free PMC article.
-
Autophagy regulates apoptosis of colorectal cancer cells based on signaling pathways.Discov Oncol. 2024 Aug 25;15(1):367. doi: 10.1007/s12672-024-01250-3. Discov Oncol. 2024. PMID: 39182013 Free PMC article. Review.
-
In situ 10-cell RNA sequencing in tissue and tumor biopsy samples.Sci Rep. 2019 Mar 20;9(1):4836. doi: 10.1038/s41598-019-41235-9. Sci Rep. 2019. PMID: 30894605 Free PMC article.
-
Ricinus communis agglutinin I leads to rapid down-regulation of VEGFR-2 and endothelial cell apoptosis in tumor blood vessels.Am J Pathol. 2010 Apr;176(4):1927-40. doi: 10.2353/ajpath.2010.090561. Epub 2010 Feb 25. Am J Pathol. 2010. PMID: 20185574 Free PMC article.
References
-
- Fend F, Specht K, Kremer M, Quintanilla-Martinez L. Laser capture microdissection in pathology. Methods Enzymol. 2002;356:196–206. - PubMed
-
- BC Decker Inc . Cancer Medicine. 5 2000. Novel Technologies for Identification of Metastasis and Invasion-Promoting Genes and Gene Products:
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

