Comparative analysis of the heat stable proteome of radicles of Medicago truncatula seeds during germination identifies late embryogenesis abundant proteins associated with desiccation tolerance
- PMID: 16461389
- PMCID: PMC1435805
- DOI: 10.1104/pp.105.074039
Comparative analysis of the heat stable proteome of radicles of Medicago truncatula seeds during germination identifies late embryogenesis abundant proteins associated with desiccation tolerance
Abstract
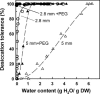
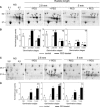
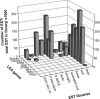
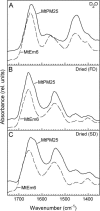
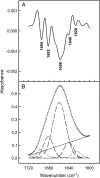
A proteomic analysis was performed on the heat stable protein fraction of imbibed radicles of Medicago truncatula seeds to investigate whether proteins can be identified that are specifically linked to desiccation tolerance (DT). Radicles were compared before and after emergence (2.8 mm long) in association with the loss of DT, and after reinduction of DT by an osmotic treatment. To separate proteins induced by the osmotic treatment from those linked with DT, the comparison was extended to 5 mm long emerged radicles for which DT could no longer be reinduced, albeit that drought tolerance was increased. The abundance of 15 polypeptides was linked with DT, out of which 11 were identified as late embryogenesis abundant proteins from different groups: MtEm6 (group 1), one isoform of DHN3 (dehydrins), MtPM25 (group 5), and three members of group 3 (MP2, an isoform of PM18, and all the isoforms of SBP65). In silico analysis revealed that their expression is likely seed specific, except for DHN3. Other isoforms of DNH3 and PM18 as well as three isoforms of the dehydrin Budcar5 were associated with drought tolerance. Changes in the abundance of MtEm6 and MtPM25 in imbibed cotyledons during the loss of DT and in developing embryos during the acquisition of DT confirmed the link of these two proteins with DT. Fourier transform infrared spectroscopy revealed that the recombinant MtPM25 and MtEm6 exhibited a certain degree of order in the hydrated state, but that they became more structured by adopting alpha helices and beta sheets during drying. A model is presented in which DT-linked late embryogenesis abundant proteins might exert different protective functions at high and low hydration levels.
Figures





Similar articles
-
MtPM25 is an atypical hydrophobic late embryogenesis-abundant protein that dissociates cold and desiccation-aggregated proteins.Plant Cell Environ. 2010 Mar;33(3):418-30. doi: 10.1111/j.1365-3040.2009.02093.x. Epub 2009 Nov 25. Plant Cell Environ. 2010. PMID: 20002332
-
Transcriptome profiling uncovers metabolic and regulatory processes occurring during the transition from desiccation-sensitive to desiccation-tolerant stages in Medicago truncatula seeds.Plant J. 2006 Sep;47(5):735-50. doi: 10.1111/j.1365-313X.2006.02822.x. Plant J. 2006. PMID: 16923015
-
LEA polypeptide profiling of recalcitrant and orthodox legume seeds reveals ABI3-regulated LEA protein abundance linked to desiccation tolerance.J Exp Bot. 2013 Nov;64(14):4559-73. doi: 10.1093/jxb/ert274. Epub 2013 Sep 16. J Exp Bot. 2013. PMID: 24043848 Free PMC article.
-
Acquisition and loss of desiccation tolerance in seeds: from experimental model to biological relevance.Planta. 2015 Mar;241(3):563-77. doi: 10.1007/s00425-014-2240-x. Epub 2015 Jan 8. Planta. 2015. PMID: 25567203 Review.
-
Proteomics of seed development, desiccation tolerance, germination and vigor.Plant Physiol Biochem. 2015 Jan;86:1-15. doi: 10.1016/j.plaphy.2014.11.003. Epub 2014 Nov 4. Plant Physiol Biochem. 2015. PMID: 25461695 Review.
Cited by
-
Role of Proteomics in Crop Stress Tolerance.Front Plant Sci. 2016 Sep 8;7:1336. doi: 10.3389/fpls.2016.01336. eCollection 2016. Front Plant Sci. 2016. PMID: 27660631 Free PMC article. Review.
-
Post-genomics studies of developmental processes in legume seeds.Plant Physiol. 2009 Nov;151(3):1023-9. doi: 10.1104/pp.109.143966. Epub 2009 Aug 12. Plant Physiol. 2009. PMID: 19675147 Free PMC article. Review. No abstract available.
-
Mimicking the plant cell interior under water stress by macromolecular crowding: disordered dehydrin proteins are highly resistant to structural collapse.Plant Physiol. 2008 Dec;148(4):1925-37. doi: 10.1104/pp.108.124099. Epub 2008 Oct 10. Plant Physiol. 2008. PMID: 18849483 Free PMC article.
-
Recent Advances in Medicago truncatula Genomics.Int J Plant Genomics. 2008;2008:256597. doi: 10.1155/2008/256597. Int J Plant Genomics. 2008. PMID: 18288239 Free PMC article.
-
An Integrated "Multi-Omics" Comparison of Embryo and Endosperm Tissue-Specific Features and Their Impact on Rice Seed Quality.Front Plant Sci. 2017 Nov 22;8:1984. doi: 10.3389/fpls.2017.01984. eCollection 2017. Front Plant Sci. 2017. PMID: 29213276 Free PMC article.
References
-
- Allison SD, Chang B, Randolph TW, Carpenter JF (1999) Hydrogen bonding between sugar and protein is responsible for inhibition of dehydration-induced protein unfolding. Arch Biochem Biophys 365: 289–298 - PubMed
-
- Alsheikh MK, Svensson JT, Randall SK (2005) Phosphorylation regulated ion-binding is a property shared by the acidic subclass of dehydrins. Plant Cell Environ 28: 1114–1122
-
- Arenas-Mena C, Raynal M, Borrell A, Varoquaux F, Cutanda MC, Stacy RAP, Pagès M, Delseny M, Culiañez-Macià FA (1999) Expression and cellular localization of Atrab 28 during Arabidopsis embryogenesis. Plant Mol Biol 40: 355–363 - PubMed
-
- Asirvatham VS, Watson BS, Sumner LW (2002) Analytical and biological variances associated with proteomic studies of Medicago truncatula by two-dimensional polyacrylamide gel electrophoresis. Proteomics 2: 960–968 - PubMed
-
- Avelange-Macherel M-H, Ly-Vu B, Delaunay J, Richomme P, Leprince O (2005) NMR metabolite profiling analysis reveals changes in phospholipid metabolism associated with the re-establishment of desiccation tolerance upon osmotic stress in germinated radicles of cucumber. Plant Cell Environ doi/10.1111/j.1365-3040.2005.01424.x - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

