Pharmacological insights obtained from structure-function studies of ionotropic glutamate receptors
- PMID: 16474411
- PMCID: PMC1760717
- DOI: 10.1038/sj.bjp.0706689
Pharmacological insights obtained from structure-function studies of ionotropic glutamate receptors
Abstract
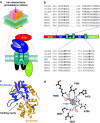
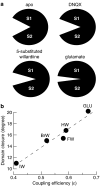
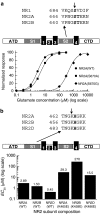
Ionotropic glutamate receptors mediate the vast majority of fast excitatory synaptic transmission in the CNS. Elucidating the structure of these proteins is central to understanding their overall function and in the last few years a tremendous amount of knowledge has been gained from the crystal structures of the ligand-binding domains of the receptor protein. These efforts have enabled us to unravel the possible mechanisms of partial agonism, agonist selectivity and desensitization. This review summarizes recent data obtained from structural studies of the binding pockets of the GluR2, GluR5/6, NR1 and NR2A subunits and discusses these studies together with homology modelling and molecular dynamics simulations that have suggested possible binding modes for full and partial agonists as well as antagonists within the binding pocket of various ionotropic glutamate receptor subunits. Comparison of the ligand-binding pockets suggests that the ligand-binding mechanisms may be conserved throughout the glutamate receptor family, although agonist selectivity may be explained by a number of features inherent to the AMPA, kainate and NMDA receptor-binding pockets such as steric occlusion, cavity size and the contribution of water-bridged interactions.
Figures




References
-
- ABELE R., SVERGUN D., KEINANEN K., KOCH M.H., MADDEN D.R. A molecular envelope of the ligand-binding domain of a glutamate receptor in the presence and absence of agonist. Biochemistry. 1999;38:10949–10957. - PubMed
-
- ARINAMINPATHY Y., SANSOM M.S.P., BIGGIN P.C. Binding site flexibility: molecular simulation of partial and full agonists within a glutamate receptor. Mol. Pharmacol. 2006;69:11–18. - PubMed
-
- ARMSTRONG N., GOUAUX E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: crystal structures of the GluR2 ligand binding core. Neuron. 2000;28:165–181. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

