A CTCF-dependent silencer located in the differentially methylated area may regulate expression of a housekeeping gene overlapping a tissue-specific gene domain
- PMID: 16478981
- PMCID: PMC1430243
- DOI: 10.1128/MCB.26.5.1589-1597.2006
A CTCF-dependent silencer located in the differentially methylated area may regulate expression of a housekeeping gene overlapping a tissue-specific gene domain
Abstract
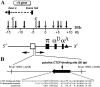
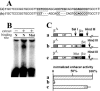
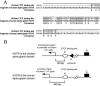
The tissue-specific chicken alpha-globin gene domain represents one of the paradigms, in terms of its constitutively open chromatin conformation and the location of several regulatory elements within the neighboring housekeeping gene. Here, we show that an 0.2-kb DNA fragment located approximately 4 kb upstream to the chicken alpha-globin gene cluster contains a binding site for the multifunctional protein factor CTCF and possesses silencer activity which depends on CTCF binding, as demonstrated by site-directed mutagenesis of the CTCF recognition sequence. CTCF was found to be associated with this recognition site in erythroid cells but not in lymphoid cells where the site is methylated. A functional promoter directing the transcription of the apparently housekeeping ggPRX gene was found 120 bp from the CTCF-dependent silencer. The data are discussed in terms of the hypothesis that the CTCF-dependent silencer stabilizes the level of ggPRX gene transcription in erythroid cells where the promoter of this gene may be influenced by positive cis-regulatory signals activating alpha-globin gene transcription.
Figures







Similar articles
-
Extensive methylation of a part of the CpG island located 3.0-4.5 kbp upstream to the chicken alpha-globin gene cluster may contribute to silencing the globin genes in non-erythroid cells.J Mol Biol. 2000 Jun 16;299(4):845-52. doi: 10.1006/jmbi.2000.3775. J Mol Biol. 2000. PMID: 10843840
-
CTCF-dependent enhancer blockers at the upstream region of the chicken alpha-globin gene domain.Nucleic Acids Res. 2004 Feb 23;32(4):1354-62. doi: 10.1093/nar/gkh301. Print 2004. Nucleic Acids Res. 2004. PMID: 14981153 Free PMC article.
-
Identification of upstream regulatory elements that repress expression of adult beta-like globin genes in a primitive erythroid environment.Blood Cells Mol Dis. 1998 Sep;24(3):356-69. doi: 10.1006/bcmd.1998.0202. Blood Cells Mol Dis. 1998. PMID: 10087993
-
CTCF function is modulated by neighboring DNA binding factors.Biochem Cell Biol. 2011 Oct;89(5):459-68. doi: 10.1139/o11-033. Epub 2011 Sep 7. Biochem Cell Biol. 2011. PMID: 21895576 Review.
-
Chromatin insulators.Annu Rev Genet. 2006;40:107-38. doi: 10.1146/annurev.genet.39.073003.113546. Annu Rev Genet. 2006. PMID: 16953792 Review.
Cited by
-
Identification of non-coding silencer elements and their regulation of gene expression.Nat Rev Mol Cell Biol. 2023 Jun;24(6):383-395. doi: 10.1038/s41580-022-00549-9. Epub 2022 Nov 7. Nat Rev Mol Cell Biol. 2023. PMID: 36344659 Review.
-
TMEM8 - a non-globin gene entrapped in the globin web.Nucleic Acids Res. 2009 Dec;37(22):7394-406. doi: 10.1093/nar/gkp838. Nucleic Acids Res. 2009. PMID: 19820109 Free PMC article.
-
Uncovering the whole genome silencers of human cells via Ss-STARR-seq.Nat Commun. 2025 Jan 16;16(1):723. doi: 10.1038/s41467-025-55852-8. Nat Commun. 2025. PMID: 39820458 Free PMC article.
-
CTCF demarcates chicken embryonic α-globin gene autonomous silencing and contributes to adult stage-specific gene expression.Epigenetics. 2013 Aug;8(8):827-38. doi: 10.4161/epi.25472. Epub 2013 Jul 3. Epigenetics. 2013. PMID: 23880533 Free PMC article.
-
Epigenetic boundaries of tumour suppressor gene promoters: the CTCF connection and its role in carcinogenesis.J Cell Mol Med. 2006 Jul-Sep;10(3):554-68. doi: 10.1111/j.1582-4934.2006.tb00420.x. J Cell Mol Med. 2006. PMID: 16989720 Free PMC article. Review.
References
-
- Bell, A. C., A. G. West, and G. Felsenfeld. 1999. The protein CTCF is required for the enhancer-blocking activity of vertebrate insulators. Cell 98:387-396. - PubMed
-
- Bell, A. C., and G. Felsenfeld. 2000. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405:482-485. - PubMed
-
- Beug, H., G. Doederlein, C. Freudenstrein, and T. Graf. 1979. Erythroblast cell lines transformed by a temperature sensitive mutant of avian erythroblastosis virus. A model system to study erythroid differentiation in vitro. J. Cell Physiol. 1:195-207. - PubMed
-
- Beug, H., S. Palmieri, C. Freudenstein, H. Zentgraf, and T. Graf. 1982. Hormone-dependent terminal differentiation in vitro of chicken erythroleukemia cells transformed by ts mutants of avian erythroblastosis virus. Cell 28:907-919. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
