Agrobacterium tumefaciens VirB8 structure reveals potential protein-protein interaction sites
- PMID: 16481621
- PMCID: PMC1413848
- DOI: 10.1073/pnas.0511216103
Agrobacterium tumefaciens VirB8 structure reveals potential protein-protein interaction sites
Abstract
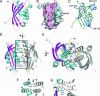
Bacterial type IV secretion systems (T4SS) translocate DNA and/or proteins to recipient cells, thus providing a mechanism for conjugative transfer of genetic material and bacterial pathogenesis. Here we describe the first structure of a core component from the archetypal Agrobacterium tumefaciens T4SS: the 2.2-A resolution crystal structure of the VirB8 periplasmic domain (pVirB8(AT)). VirB8 forms a dimer in the crystal, and we identify residues likely important for stabilization of the dimer interface. Structural comparison of pVirB8(AT) with Brucella suis VirB8 confirms that the monomers have a similar fold. In addition, the pVirB8(AT) dimer superimposes very closely on the B. suis VirB8 dimer, supporting the proposal that dimer formation in the crystal reflects self-interactions that are biologically significant. The evolutionary conservation level for each residue was obtained from a data set of 84 VirB8 homologs and projected onto the protein structure to indicate conserved surface patches that likely contact other T4SS proteins.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures


Similar articles
-
Structural Analysis and Inhibition of TraE from the pKM101 Type IV Secretion System.J Biol Chem. 2016 Nov 4;291(45):23817-23829. doi: 10.1074/jbc.M116.753327. Epub 2016 Sep 15. J Biol Chem. 2016. PMID: 27634044 Free PMC article.
-
The dimer interface of Agrobacterium tumefaciens VirB8 is important for type IV secretion system function, stability, and association of VirB2 with the core complex.J Bacteriol. 2011 May;193(9):2097-106. doi: 10.1128/JB.00907-10. Epub 2011 Mar 11. J Bacteriol. 2011. PMID: 21398549 Free PMC article.
-
Interactions between Brucella suis VirB8 and its homolog TraJ from the plasmid pSB102 underline the dynamic nature of type IV secretion systems.J Bacteriol. 2009 May;191(9):2985-92. doi: 10.1128/JB.01426-08. Epub 2009 Feb 27. J Bacteriol. 2009. PMID: 19251859 Free PMC article.
-
VirB8: a conserved type IV secretion system assembly factor and drug target.Biochem Cell Biol. 2006 Dec;84(6):890-9. doi: 10.1139/o06-148. Biochem Cell Biol. 2006. PMID: 17215876 Review.
-
The Agrobacterium VirB/VirD4 T4SS: Mechanism and Architecture Defined Through In Vivo Mutagenesis and Chimeric Systems.Curr Top Microbiol Immunol. 2018;418:233-260. doi: 10.1007/82_2018_94. Curr Top Microbiol Immunol. 2018. PMID: 29808338 Free PMC article. Review.
Cited by
-
VirB8-like protein TraH is crucial for DNA transfer in Enterococcus faecalis.Sci Rep. 2016 Apr 22;6:24643. doi: 10.1038/srep24643. Sci Rep. 2016. PMID: 27103580 Free PMC article.
-
Orientia tsutsugamushi: comprehensive analysis of the mobilome of a highly fragmented and repetitive genome reveals the capacity for ongoing lateral gene transfer in an obligate intracellular bacterium.mSphere. 2023 Dec 20;8(6):e0026823. doi: 10.1128/msphere.00268-23. Epub 2023 Oct 18. mSphere. 2023. PMID: 37850800 Free PMC article.
-
Questing functions and structures of hypothetical proteins from Campylobacter jejuni: a computer-aided approach.Biosci Rep. 2020 Jun 26;40(6):BSR20193939. doi: 10.1042/BSR20193939. Biosci Rep. 2020. PMID: 32458979 Free PMC article.
-
Biological diversity of prokaryotic type IV secretion systems.Microbiol Mol Biol Rev. 2009 Dec;73(4):775-808. doi: 10.1128/MMBR.00023-09. Microbiol Mol Biol Rev. 2009. PMID: 19946141 Free PMC article. Review.
-
Inner membrane components of the plasmid pKM101 type IV secretion system TraE and TraD are DNA-binding proteins.Sci Rep. 2025 Mar 4;15(1):7530. doi: 10.1038/s41598-025-85446-9. Sci Rep. 2025. PMID: 40032854 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

