Use of thymine limitation and thymine starvation to study bacterial physiology and cytology
- PMID: 16484178
- PMCID: PMC1426543
- DOI: 10.1128/JB.188.5.1667-1679.2006
Use of thymine limitation and thymine starvation to study bacterial physiology and cytology
Figures

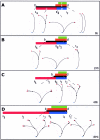
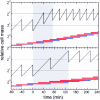

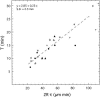
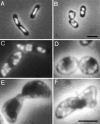
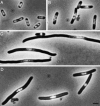
References
-
- Aarsman, M. E. G., A. Piette, C. Fraipont, T. M. F. Vinkelvleugel, M. Nguyen-Distèche, M., and T. den Blaauwen. 2005. Maturation of the Escherichia coli divisome occurs in two steps. Mol. Microbiol. 55:1631-1645. - PubMed
-
- Ahmad, S. I., S. H. Kirk, and A. Eisenstark. 1998. Thymine metabolism and thymineless death in prokaryotes and eukaryotes. Annu. Rev. Microbiol. 52:591-625. - PubMed
-
- Bazill, G. W. 1967. Lethal unbalanced growth in bacteria. Nature 216:346-349. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

