Species- and strain-specific control of a complex, flexible regulon by Bordetella BvgAS
- PMID: 16484188
- PMCID: PMC1426559
- DOI: 10.1128/JB.188.5.1775-1785.2006
Species- and strain-specific control of a complex, flexible regulon by Bordetella BvgAS
Abstract
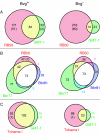
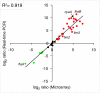
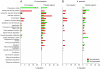
The Bordetella master virulence regulatory system, BvgAS, controls a spectrum of gene expression states, including the virulent Bvg(+) phase, the avirulent Bvg(-) phase, and at least one Bvg-intermediate (Bvg(i)) phase. We set out to define the species- and strain-specific features of this regulon based on global gene expression profiling. Rather than functioning as a switch, Bvg controls a remarkable continuum of gene expression states, with hundreds of genes maximally expressed in intermediate phases between the Bvg(+) and Bvg(-) poles. Comparative analysis of Bvg regulation in B. pertussis and B. bronchiseptica revealed a relatively conserved Bvg(+) phase transcriptional program and identified previously uncharacterized candidate virulence factors. In contrast, control of Bvg(-)- and Bvg(i)-phase genes diverged substantially between species; regulation of metabolic, transporter, and motility loci indicated an increased capacity in B. bronchiseptica, compared to B. pertussis, for ex vivo adaptation. Strain comparisons also demonstrated variation in gene expression patterns within species. Among the genes with the greatest variability in patterns of expression, predicted promoter sequences were nearly identical. Our data suggest that the complement of transcriptional regulators is largely responsible for transcriptional diversity. In support of this hypothesis, many putative transcriptional regulators that were Bvg regulated in B. bronchiseptica were deleted, inactivated, or unregulated by BvgAS in B. pertussis. We propose the concept of a "flexible regulon." This flexible regulon may prove to be important for pathogen evolution and the diversification of host range specificity.
Figures



References
-
- Akerley, B. J., P. A. Cotter, and J. F. Miller. 1995. Ectopic expression of the flagellar regulon alters development of the Bordetella-host interaction. Cell 80:611-620. - PubMed
-
- Bailey, T. L., and C. Elkan. 1994. Fitting a mixture model by expectation maximization to discover motifs in biopolymers, p. 28-36, In Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology. AAAI Press, Menlo Park, Calif. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

