Gene products required for de novo synthesis of polysialic acid in Escherichia coli K1
- PMID: 16484189
- PMCID: PMC1426546
- DOI: 10.1128/JB.188.5.1786-1797.2006
Gene products required for de novo synthesis of polysialic acid in Escherichia coli K1
Abstract
Escherichia coli K1 is responsible for 80% of E. coli neonatal meningitis and is a common pathogen in urinary tract infections. Bacteria of this serotype are encapsulated with the alpha(2-8)-polysialic acid NeuNAc(alpha2-8), common to several bacterial pathogens. The gene cluster encoding the pathway for synthesis of this polymer is organized into three regions: (i) kpsSCUDEF, (ii) neuDBACES, and (iii) kpsMT. The K1 polysialyltransferase, NeuS, cannot synthesize polysialic acid de novo without other products of the gene cluster. Membranes isolated from strains having the entire K1 gene cluster can synthesize polysialic acid de novo. We designed a series of plasmid constructs containing fragments of regions 1 and 2 in two compatible vectors to determine the minimum number of gene products required for de novo synthesis of the polysialic acid from CMP-NeuNAc in K1 E. coli. We measured the ability of the various combinations of region 1 and 2 fragments to restore polysialyltransferase activity in vitro in the absence of exogenously added polysaccharide acceptor. The products of region 2 genes neuDBACES alone were not sufficient to support de novo synthesis of polysialic acid in vitro. Only membrane fractions harboring NeuES and KpsCS could form sialic polymer in the absence of exogenous acceptor at the concentrations formed by wild-type E. coli K1 membranes. Membrane fractions harboring NeuES and KpsC together could form small quantities of the sialic polymer de novo.
Figures




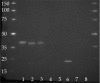
References
-
- Adlam, C., J. M. Knights, A. Mugridge, J. M. Williams, and J. C. Lindon. 1987. Production of colominic acid by Pasteurella haemolytica serotype A2 organisms. FEMS Microbiol. Lett. 42:23-25.
-
- Bøvre, K., K. Bryn, O. Closs, N. Hagen, and L. O. Frøholm. 1983. Surface polysaccharide of Moraxella non-liquefaciens identical to Neisseria meningitidis group B capsular polysaccharide. A chemical and immunological investigation. NIPH Ann. 6:65-73. - PubMed
-
- Daines, D. A., L. F. Wright, D. O. Chaffin, C. E. Rubens, and R. P. Silver. 2000. NeuD plays a role in the synthesis of sialic acid in Escherichia coli K1. FEMS Microbiol. Lett. 189:281-284. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

