The Arabidopsis Aux/IAA protein family has diversified in degradation and auxin responsiveness
- PMID: 16489122
- PMCID: PMC1383644
- DOI: 10.1105/tpc.105.039172
The Arabidopsis Aux/IAA protein family has diversified in degradation and auxin responsiveness
Abstract
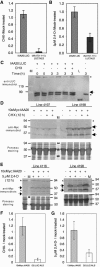
Rapid, auxin-responsive degradation of multiple auxin/indole-3-acetic acid (Aux/IAA) proteins is essential for plant growth and development. Domain II residues were previously shown to be required for the degradation of several Arabidopsis thaliana Aux/IAA proteins. We examined the degradation of additional full-length family members and the proteolytic importance of N-terminal residues outside domain II using luciferase (LUC) fusions. Elimination of domain I did not affect degradation. However, substituting an Arg for a conserved Lys between domains I and II specifically impaired basal degradation without compromising the auxin-mediated acceleration of degradation. IAA8, IAA9, and IAA28 contain domain II and a conserved Lys, but they were degraded more slowly than previously characterized family members when expressed as LUC fusions, suggesting that sequences outside domain II influence proteolysis. We analyzed the degradation of IAA31, with a region somewhat similar to domain II but without the conserved Lys, and of IAA20, which lacks domain II and the conserved Lys. Both IAA20:LUC and epitope-tagged IAA20 were long-lived, and their longevity was not influenced by auxin. Epitope-tagged IAA31 was long-lived, like IAA20, but by contrast, it showed accelerated degradation in response to auxin. The existence of long-lived and auxin-insensitive Aux/IAA proteins suggeststhat they may play a novel role in auxin signaling.
Figures




References
-
- Abel, S., Nguyen, M.D., and Theologis, A. (1995). The PSIAA4/5-like family of early auxin-inducible mRNAs in Arabidopsis thaliana. J. Mol. Biol. 251 533–549. - PubMed
-
- Abel, S., and Theologis, A. (1995). A polymorphic bipartite motif signals nuclear targeting of early auxin-inducible proteins related to PSIAA4 from pea (Pisum sativum). Plant J. 8 87–96. - PubMed
-
- Balint, E., Bates, S., and Vousden, K.H. (1999). Mdm2 binds p73 alpha without targeting degradation. Oncogene 18 3923–3929. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

