Combined top-down and bottom-up proteomics identifies a phosphorylation site in stem-loop-binding proteins that contributes to high-affinity RNA binding
- PMID: 16492733
- PMCID: PMC1413926
- DOI: 10.1073/pnas.0511289103
Combined top-down and bottom-up proteomics identifies a phosphorylation site in stem-loop-binding proteins that contributes to high-affinity RNA binding
Abstract
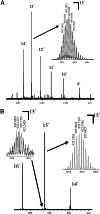
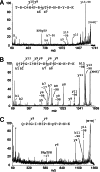
The stem-loop-binding protein (SLBP) is involved in multiple aspects of histone mRNA metabolism. To characterize the modification status and sites of SLBP, we combined mass spectrometric bottom-up (analysis of peptides) and top-down (analysis of intact proteins) proteomic approaches. Drosophilia SLBP is heavily phosphorylated, containing up to seven phosphoryl groups. Accurate M(r) determination by Fourier transform ion cyclotron resonance (FTICR)-MS and FTICR-MS top-down experiments using a variety of dissociation techniques show there is removal of the initiator methionine and acetylation of the N terminus in the baculovirus-expressed protein, and that T230 is stoichiometrically phosphorylated. T230 is highly conserved; we have determined that this site is also completely phosphorylated in baculovirus-expressed mammalian SLBP and extensively phosphorylated in both Drosophila and mammalian cultured cells. Removal of the phosphoryl group from T230 by either dephosphorylation or mutation results in a 7-fold reduction in the affinity of SLBP for the stem-loop RNA.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

