Memory T and memory B cells share a transcriptional program of self-renewal with long-term hematopoietic stem cells
- PMID: 16492737
- PMCID: PMC1413911
- DOI: 10.1073/pnas.0511137103
Memory T and memory B cells share a transcriptional program of self-renewal with long-term hematopoietic stem cells
Abstract
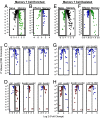
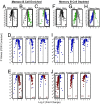
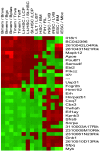
The only cells of the hematopoietic system that undergo self-renewal for the lifetime of the organism are long-term hematopoietic stem cells and memory T and B cells. To determine whether there is a shared transcriptional program among these self-renewing populations, we first compared the gene-expression profiles of naïve, effector and memory CD8(+) T cells with those of long-term hematopoietic stem cells, short-term hematopoietic stem cells, and lineage-committed progenitors. Transcripts augmented in memory CD8(+) T cells relative to naïve and effector T cells were selectively enriched in long-term hematopoietic stem cells and were progressively lost in their short-term and lineage-committed counterparts. Furthermore, transcripts selectively decreased in memory CD8(+) T cells were selectively down-regulated in long-term hematopoietic stem cells and progressively increased with differentiation. To confirm that this pattern was a general property of immunologic memory, we turned to independently generated gene expression profiles of memory, naïve, germinal center, and plasma B cells. Once again, memory-enriched and -depleted transcripts were also appropriately augmented and diminished in long-term hematopoietic stem cells, and their expression correlated with progressive loss of self-renewal function. Thus, there appears to be a common signature of both up- and down-regulated transcripts shared between memory T cells, memory B cells, and long-term hematopoietic stem cells. This signature was not consistently enriched in neural or embryonic stem cell populations and, therefore, appears to be restricted to the hematopoeitic system. These observations provide evidence that the shared phenotype of self-renewal in the hematopoietic system is linked at the molecular level.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures

References
-
- Fearon D. T., Manders P., Wagner S. D. Science. 2001;293:248–250. - PubMed
-
- Kaech S. M., Wherry E. J., Ahmed R. Nat. Rev. Immunol. 2002;2:251–262. - PubMed
-
- Lanzavecchia A., Sallusto F. Nat. Rev. Immunol. 2002;2:982–987. - PubMed
-
- Kondo M., Wagers A. J., Manz M. G., Prohaska S. S., Scherer D. C., Beilhack G. F., Shizuru J. A., Weissman I. L. Annu. Rev. Immunol. 2002:759–806. - PubMed
-
- Schluns K. S., Lefrancois L. Nat. Rev. Immunol. 2003;3:269–279. - PubMed
Publication types
MeSH terms
Grants and funding
- T32 HL0762-18/HL/NHLBI NIH HHS/United States
- K08 AI063386/AI/NIAID NIH HHS/United States
- P01 DK053074/DK/NIDDK NIH HHS/United States
- 2 P30 DK36836-17/DK/NIDDK NIH HHS/United States
- P01DK053074/DK/NIDDK NIH HHS/United States
- R01HL058770/HL/NHLBI NIH HHS/United States
- R37 AI051530/AI/NIAID NIH HHS/United States
- T32 DK007260/DK/NIDDK NIH HHS/United States
- P30 DK036836/DK/NIDDK NIH HHS/United States
- 5 T32 DK007260-24/DK/NIDDK NIH HHS/United States
- R01 AI051530/AI/NIAID NIH HHS/United States
- R01AI047457/AI/NIAID NIH HHS/United States
- R01 AI047457/AI/NIAID NIH HHS/United States
- AI51530-01/AI/NIAID NIH HHS/United States
- R01 HL058770/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

