Comparative study of the effects of heptameric slippery site composition on -1 frameshifting among different eukaryotic systems
- PMID: 16497657
- PMCID: PMC1421095
- DOI: 10.1261/rna.2225206
Comparative study of the effects of heptameric slippery site composition on -1 frameshifting among different eukaryotic systems
Abstract
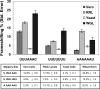
Studies of programmed -1 ribosomal frameshifting (-1 PRF) have been approached over the past two decades by many different laboratories using a diverse array of virus-derived frameshift signals in translational assay systems derived from a variety of sources. Though it is generally acknowledged that both absolute and relative -1 PRF efficiency can vary in an assay system-dependent manner, no methodical study of this phenomenon has been undertaken. To address this issue, a series of slippery site mutants of the SARS-associated coronavirus frameshift signal were systematically assayed in four different eukaryotic translational systems. HIV-1 promoted frameshifting was also compared between Escherichia coli and a human T-cell line expression systems. The results of these analyses highlight different aspects of each system, suggesting in general that (1) differences can be due to the assay systems themselves; (2) phylogenetic differences in ribosome structure can affect frameshifting efficiency; and (3) care must be taken to employ the closest phylogenetic match between a specific -1 PRF signal and the choice of translational assay system.
Figures
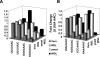
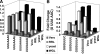
References
-
- Barak, Z., Gallant, J., Lindsley, D., Kwieciszewki, B., and Heidel, D. 1996. Enhanced ribosome frameshifting in stationary phase cells. J. Mol. Biol. 263: 140–148. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
