Activation energies for dissociation of double strand oligonucleotide anions: evidence for watson-crick base pairing in vacuo
- PMID: 16498487
- PMCID: PMC1380309
- DOI: 10.1021/ja973534h
Activation energies for dissociation of double strand oligonucleotide anions: evidence for watson-crick base pairing in vacuo
Abstract
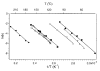
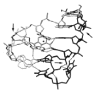
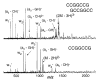
The dissociation kinetics of a series of complementary and noncomplementary DNA duplexes, (TGCA)(2) (3-), (CCGG)(2) (3-), (AATTAAT)(2) (3-), (CCGGCCG)(2) (3-), A(7)*T(7) (3-), A(7)*A(7) (3-), T(7)*T(7) (3-), and A(7)*C(7) (3-) were investigated using blackbody infrared radiative dissociation in a Fourier transform mass spectrometer. From the temperature dependence of the unimolecular dissociation rate constants, Arrhenius activation parameters in the zero-pressure limit are obtained. Activation energies range from 1.2 to 1.7 eV, and preexponential factors range from 10(13) to 10(19) s(-1). Dissociation of the duplexes results in cleavage of the noncovalent bonds and/or cleavage of covalent bonds leading to loss of a neutral nucleobase followed by backbone cleavage producing sequence-specific (a - base) and w ions. Four pieces of evidence are presented which indicate that Watson-Crick (WC) base pairing is preserved in complementary DNA duplexes in the gas phase: i. the activation energy for dissociation of the complementary dimer, A(7)*T(7) (3-), to the single strands is significantly higher than that for the related noncomplementary A(7)*A(7) (3-) and T(7)*T(7) (3-) dimers, indicating a stronger interaction between strands with a specific base sequence, ii. extensive loss of neutral adenine occurs for A(7)*A(7) (3-) and A(7)*C(7) (3-) but not for A(7)*T(7) (3-) consistent with this process being shut down by WC hydrogen bonding, iii. a correlation is observed between the measured activation energy for dissociation to single strands and the dimerization enthalpy (-DeltaH(d)) in solution, and iv. molecular dynamics carried out at 300 and 400 K indicate that WC base pairing is preserved for A(7)*T(7) (3-) duplex, although the helical structure is essentially lost. In combination, these results provide strong evidence that WC base pairing can exist in the complete absence of solvent.
Figures

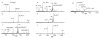

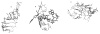


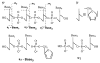
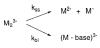
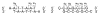
Similar articles
-
Blackbody infrared radiative dissociation of oligonucleotide anions.J Am Soc Mass Spectrom. 1998 Nov;9(11):1117-24. doi: 10.1016/S1044-0305(98)00098-1. J Am Soc Mass Spectrom. 1998. PMID: 9794082 Free PMC article.
-
Thermal decomposition of multiply charged T-rich oligonucleotide anions in the gas phase. Influence of internal solvation on the arrhenius parameters for neutral base loss.J Am Soc Mass Spectrom. 2006 Sep;17(9):1229-38. doi: 10.1016/j.jasms.2006.05.002. Epub 2006 Jun 16. J Am Soc Mass Spectrom. 2006. PMID: 16782356
-
Effects of charge state on fragmentation pathways, dynamics, and activation energies of ubiquitin ions measured by blackbody infrared radiative dissociation.Anal Chem. 1997 Mar 15;69(6):1119-26. doi: 10.1021/ac960804q. Anal Chem. 1997. PMID: 9075403 Free PMC article.
-
Dissociation energies of deoxyribose nucleotide dimer anions measured using blackbody infrared radiative dissociation.J Am Soc Mass Spectrom. 1999 Nov;10(11):1095-104. doi: 10.1016/S1044-0305(99)00087-2. J Am Soc Mass Spectrom. 1999. PMID: 10536816 Free PMC article.
-
Fingerprints of bonding motifs in DNA duplexes of adenine and thymine revealed from circular dichroism: synchrotron radiation experiments and TDDFT calculations.J Phys Chem B. 2009 Jul 16;113(28):9614-9. doi: 10.1021/jp9032029. J Phys Chem B. 2009. PMID: 19537699
Cited by
-
The Contribution of the Activation Entropy to the Gas-Phase Stability of Modified Nucleic Acid Duplexes.J Am Soc Mass Spectrom. 2016 Jul;27(7):1186-96. doi: 10.1007/s13361-016-1391-3. Epub 2016 Apr 14. J Am Soc Mass Spectrom. 2016. PMID: 27080005
-
Further studies on the origins of asymmetric charge partitioning in protein homodimers.J Am Soc Mass Spectrom. 2004 Oct;15(10):1408-1415. doi: 10.1016/j.jasms.2004.06.006. J Am Soc Mass Spectrom. 2004. PMID: 15465353 Free PMC article.
-
Secondary structural characterization of oligonucleotide strands using electrospray ionization mass spectrometry.Nucleic Acids Res. 2005 Jun 28;33(11):3659-66. doi: 10.1093/nar/gki671. Print 2005. Nucleic Acids Res. 2005. PMID: 15985683 Free PMC article.
-
A role for the MS analysis of nucleic acids in the post-genomics age.J Am Soc Mass Spectrom. 2010 Jan;21(1):1-13. doi: 10.1016/j.jasms.2009.09.006. Epub 2009 Sep 17. J Am Soc Mass Spectrom. 2010. PMID: 19897384
-
Tandem mass spectrometry of protein-protein complexes: cytochrome c-cytochrome b5.J Am Soc Mass Spectrom. 2002 Jan;13(1):59-71. doi: 10.1016/S1044-0305(01)00331-2. J Am Soc Mass Spectrom. 2002. PMID: 11777200
References
-
- Fenn JB, Mann M, Meng CK, Wong SF, Whitehouse CM. Science. 1989;246:64–71. - PubMed
-
- Hillenkamp F, Karas M, Beavis RC, Chait BT. Anal Chem. 1991;63:1193A–1202A. - PubMed
-
- Bowers MT, Marshall AG, McLafferty FW. J Phys Chem. 1996;100:12897–12910.
-
- McLafferty FW. Acc Chem Res. 1994;27:379–386. and references therein.
- Kelleher NL, Senko MW, Siegel MM, McLafferty FW. J Am Soc Mass Spectrom. 1997;8:380–383.
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
