Characterization of 43 non-protein-coding mRNA genes in Arabidopsis, including the MIR162a-derived transcripts
- PMID: 16500993
- PMCID: PMC1435803
- DOI: 10.1104/pp.105.073817
Characterization of 43 non-protein-coding mRNA genes in Arabidopsis, including the MIR162a-derived transcripts
Abstract
Messenger RNAs that do not contain a long open reading frame (ORF) or non-protein-coding RNAs (npcRNAs) are an emerging novel class of transcripts. Their functions may involve the RNA molecule itself and/or short ORF-encoded peptides. npcRNA genes are difficult to identify using standard gene prediction programs that rely on the presence of relatively long ORFs. Here, we used detailed bioinformatic analyses of expressed sequence tag/cDNA databases to detect a restricted set of npcRNAs in the Arabidopsis (Arabidopsis thaliana) genome and further characterized these transcripts using a combination of bioinformatic and molecular approaches. Compositional analyses revealed strong nucleotide strand asymmetries in the npcRNAs, as well as a biased GC content, suggesting the existence of functional constraints on these RNAs. Thirteen of these transcripts display tissue-specific expression patterns, and three are regulated in conditions affecting root architecture. The npcRNA 78 gene contains the miR162 sequence in an alternative intron and corresponds to the MIR162a locus. Although DICER-LIKE 1 (DCL1) mRNA is known to be regulated by miR162-guided cleavage, its level does not change in a mir162a mutant. Alternative splicing of npcRNA 78 leads to several transcript isoforms, which all accumulate in a dcl1 mutant. This suggests that npcRNA 78 is a genuine substrate of DCL1 and that splicing of this microRNA primary transcript and miR162 processing are competitive nuclear events. Our results provide new insights into Arabidopsis npcRNA biology and the potential roles of these genes.
Figures




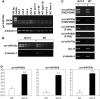
Similar articles
-
Riboregulators in plant development.Biochem Soc Trans. 2007 Dec;35(Pt 6):1638-42. doi: 10.1042/BST0351638. Biochem Soc Trans. 2007. PMID: 18031282 Review.
-
Arabidopsis small RNAs and their targets during cyst nematode parasitism.Mol Plant Microbe Interact. 2008 Dec;21(12):1622-34. doi: 10.1094/MPMI-21-12-1622. Mol Plant Microbe Interact. 2008. PMID: 18986258
-
Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses.Genome Res. 2009 Jan;19(1):57-69. doi: 10.1101/gr.080275.108. Epub 2008 Nov 7. Genome Res. 2009. PMID: 18997003 Free PMC article.
-
Computational detection of microRNAs targeting transcription factor genes in Arabidopsis thaliana.Comput Biol Chem. 2005 Oct;29(5):360-7. doi: 10.1016/j.compbiolchem.2005.08.005. Epub 2005 Oct 10. Comput Biol Chem. 2005. PMID: 16221572
-
Dual RNAs in plants.Biochimie. 2011 Nov;93(11):1950-4. doi: 10.1016/j.biochi.2011.07.028. Epub 2011 Jul 31. Biochimie. 2011. PMID: 21824505 Review.
Cited by
-
Protein-coding cis-natural antisense transcripts have high and broad expression in Arabidopsis.Plant Physiol. 2013 Apr;161(4):2171-80. doi: 10.1104/pp.112.212100. Epub 2013 Mar 1. Plant Physiol. 2013. PMID: 23457227 Free PMC article.
-
Deciphering the plant splicing code: experimental and computational approaches for predicting alternative splicing and splicing regulatory elements.Front Plant Sci. 2012 Feb 7;3:18. doi: 10.3389/fpls.2012.00018. eCollection 2012. Front Plant Sci. 2012. PMID: 22645572 Free PMC article.
-
Overexpression of microRNA395c or 395e affects differently the seed germination of Arabidopsis thaliana under stress conditions.Planta. 2010 Nov;232(6):1447-54. doi: 10.1007/s00425-010-1267-x. Epub 2010 Sep 14. Planta. 2010. PMID: 20839006
-
Bidirectional processing of pri-miRNAs with branched terminal loops by Arabidopsis Dicer-like1.Nat Struct Mol Biol. 2013 Sep;20(9):1106-15. doi: 10.1038/nsmb.2646. Epub 2013 Aug 11. Nat Struct Mol Biol. 2013. PMID: 23934148 Free PMC article.
-
Linking discoveries, mechanisms, and technologies to develop a clearer perspective on plant long noncoding RNAs.Plant Cell. 2023 May 29;35(6):1762-1786. doi: 10.1093/plcell/koad027. Plant Cell. 2023. PMID: 36738093 Free PMC article. Review.
References
-
- Allen E, Xie Z, Gustafson AM, Carrington JC (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121: 207–221 - PubMed
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657 - PubMed
-
- Baker CC, Sieber P, Wellmer F, Meyerowitz EM (2005) The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr Biol 15: 303–315 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

