The effect of oligonucleotide microarray data pre-processing on the analysis of patient-cohort studies
- PMID: 16512908
- PMCID: PMC1481623
- DOI: 10.1186/1471-2105-7-105
The effect of oligonucleotide microarray data pre-processing on the analysis of patient-cohort studies
Abstract
Background: Intensity values measured by Affymetrix microarrays have to be both normalized, to be able to compare different microarrays by removing non-biological variation, and summarized, generating the final probe set expression values. Various pre-processing techniques, such as dChip, GCRMA, RMA and MAS have been developed for this purpose. This study assesses the effect of applying different pre-processing methods on the results of analyses of large Affymetrix datasets. By focusing on practical applications of microarray-based research, this study provides insight into the relevance of pre-processing procedures to biology-oriented researchers.
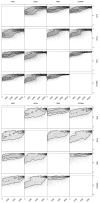
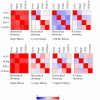
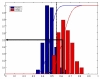
Results: Using two publicly available datasets, i.e., gene-expression data of 285 patients with Acute Myeloid Leukemia (AML, Affymetrix HG-U133A GeneChip) and 42 samples of tumor tissue of the embryonal central nervous system (CNS, Affymetrix HuGeneFL GeneChip), we tested the effect of the four pre-processing strategies mentioned above, on (1) expression level measurements, (2) detection of differential expression, (3) cluster analysis and (4) classification of samples. In most cases, the effect of pre-processing is relatively small compared to other choices made in an analysis for the AML dataset, but has a more profound effect on the outcome of the CNS dataset. Analyses on individual probe sets, such as testing for differential expression, are affected most; supervised, multivariate analyses such as classification are far less sensitive to pre-processing.
Conclusion: Using two experimental datasets, we show that the choice of pre-processing method is of relatively minor influence on the final analysis outcome of large microarray studies whereas it can have important effects on the results of a smaller study. The data source (platform, tissue homogeneity, RNA quality) is potentially of bigger importance than the choice of pre-processing method.
Figures
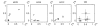
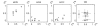
Similar articles
-
A comprehensive evaluation of multicategory classification methods for microarray gene expression cancer diagnosis.Bioinformatics. 2005 Mar 1;21(5):631-43. doi: 10.1093/bioinformatics/bti033. Epub 2004 Sep 16. Bioinformatics. 2005. PMID: 15374862
-
Interactively optimizing signal-to-noise ratios in expression profiling: project-specific algorithm selection and detection p-value weighting in Affymetrix microarrays.Bioinformatics. 2004 Nov 1;20(16):2534-44. doi: 10.1093/bioinformatics/bth280. Epub 2004 Apr 29. Bioinformatics. 2004. PMID: 15117752
-
SplicerAV: a tool for mining microarray expression data for changes in RNA processing.BMC Bioinformatics. 2010 Feb 25;11:108. doi: 10.1186/1471-2105-11-108. BMC Bioinformatics. 2010. PMID: 20184770 Free PMC article.
-
Clinical uses of microarrays in cancer research.Methods Mol Med. 2008;141:87-113. doi: 10.1007/978-1-60327-148-6_6. Methods Mol Med. 2008. PMID: 18453086 Free PMC article. Review.
-
Tissue microarrays in clinical oncology.Semin Radiat Oncol. 2008 Apr;18(2):89-97. doi: 10.1016/j.semradonc.2007.10.006. Semin Radiat Oncol. 2008. PMID: 18314063 Free PMC article. Review.
Cited by
-
Prediction of early breast cancer patient survival using ensembles of hypoxia signatures.PLoS One. 2018 Sep 14;13(9):e0204123. doi: 10.1371/journal.pone.0204123. eCollection 2018. PLoS One. 2018. PMID: 30216362 Free PMC article.
-
Comparative analysis of methods for gene transcription profiling data derived from different microarray technologies in rat and mouse models of diabetes.BMC Genomics. 2009 Feb 5;10:63. doi: 10.1186/1471-2164-10-63. BMC Genomics. 2009. PMID: 19196459 Free PMC article.
-
Unifying gene expression measures from multiple platforms using factor analysis.PLoS One. 2011 Mar 11;6(3):e17691. doi: 10.1371/journal.pone.0017691. PLoS One. 2011. PMID: 21436879 Free PMC article.
-
Ensemble analyses improve signatures of tumour hypoxia and reveal inter-platform differences.BMC Bioinformatics. 2014 Jun 6;15:170. doi: 10.1186/1471-2105-15-170. BMC Bioinformatics. 2014. PMID: 24902696 Free PMC article.
-
Statistical analysis of an RNA titration series evaluates microarray precision and sensitivity on a whole-array basis.BMC Bioinformatics. 2006 Nov 22;7:511. doi: 10.1186/1471-2105-7-511. BMC Bioinformatics. 2006. PMID: 17118209 Free PMC article.
References
-
- Affymetrix Microarray Suite User Guide. 2001.
-
- Naef F, Magnasco MO. Solving the riddle of the bright mismatches: labeling and effective binding in oligonucleotide arrays. Phys Rev E Stat Nonlin Soft Matter Phys. 2003;68:11906. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

