Bayesian analyses of multiple epistatic QTL models for body weight and body composition in mice
- PMID: 16545150
- PMCID: PMC5002393
- DOI: 10.1017/S0016672306007944
Bayesian analyses of multiple epistatic QTL models for body weight and body composition in mice
Abstract
To comprehensively investigate the genetic architecture of growth and obesity, we performed Bayesian analyses of multiple epistatic quantitative trait locus (QTL) models for body weights at five ages (12 days, 3, 6, 9 and 12 weeks) and body composition traits (weights of two fat pads and five organs) in mice produced from a cross of the F1 between M16i (selected for rapid growth rate) and CAST/Ei (wild-derived strain of small and lean mice) back to M16i. Bayesian model selection revealed a temporally regulated network of multiple QTL for body weight, involving both strong main effects and epistatic effects. No QTL had strong support for both early and late growth, although overlapping combinations of main and epistatic effects were observed at adjacent ages. Most main effects and epistatic interactions had an opposite effect on early and late growth. The contribution of epistasis was more pronounced for body weights at older ages. Body composition traits were also influenced by an interacting network of multiple QTLs. Several main and epistatic effects were shared by the body composition and body weight traits, suggesting that pleiotropy plays an important role in growth and obesity.
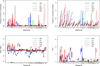
Figures
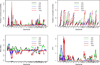
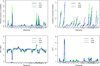
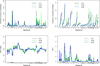
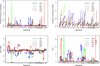
Similar articles
-
Genetic influences on growth and body composition in mice: multilocus interactions.Int J Obes (Lond). 2009 Jan;33(1):89-95. doi: 10.1038/ijo.2008.215. Epub 2008 Nov 4. Int J Obes (Lond). 2009. PMID: 18982013 Free PMC article.
-
The contribution of epistatic pleiotropy to the genetic architecture of covariation among polygenic traits in mice.Evol Dev. 2006 Sep-Oct;8(5):468-76. doi: 10.1111/j.1525-142X.2006.00120.x. Evol Dev. 2006. PMID: 16925682
-
QTLs for pre- and postweaning body weight and body composition in selected mice.Mamm Genome. 2004 Aug;15(8):593-609. doi: 10.1007/s00335-004-3026-4. Mamm Genome. 2004. PMID: 15457339
-
QTL-based evidence for the role of epistasis in evolution.Genet Res. 2005 Oct;86(2):89-95. doi: 10.1017/S0016672305007780. Genet Res. 2005. PMID: 16356282 Review.
-
A pleiotropic-epistatic entangelement model of drug response.Drug Discov Today. 2023 Nov;28(11):103790. doi: 10.1016/j.drudis.2023.103790. Epub 2023 Sep 26. Drug Discov Today. 2023. PMID: 37758020 Review.
Cited by
-
Hierarchical generalized linear models for multiple quantitative trait locus mapping.Genetics. 2009 Mar;181(3):1101-13. doi: 10.1534/genetics.108.099556. Epub 2009 Jan 12. Genetics. 2009. PMID: 19139143 Free PMC article.
-
Extensive epistasis for olfactory behaviour, sleep and waking activity in Drosophila melanogaster.Genet Res (Camb). 2012 Feb;94(1):9-20. doi: 10.1017/S001667231200002X. Genet Res (Camb). 2012. PMID: 22353245 Free PMC article.
-
Identification of quantitative trait loci affecting body composition in a mouse intercross.Proc Natl Acad Sci U S A. 2006 Dec 26;103(52):19860-5. doi: 10.1073/pnas.0609232103. Epub 2006 Dec 18. Proc Natl Acad Sci U S A. 2006. PMID: 17179051 Free PMC article.
-
Genetic influences on growth and body composition in mice: multilocus interactions.Int J Obes (Lond). 2009 Jan;33(1):89-95. doi: 10.1038/ijo.2008.215. Epub 2008 Nov 4. Int J Obes (Lond). 2009. PMID: 18982013 Free PMC article.
-
Genomewide analysis of epistatic effects for quantitative traits in barley.Genetics. 2007 Apr;175(4):1955-63. doi: 10.1534/genetics.106.066571. Epub 2007 Feb 4. Genetics. 2007. PMID: 17277367 Free PMC article.
References
-
- Allan MF, Eisen EJ, Pomp D. The M16 Mouse: An outbred animal model of early onset obesity and diabesity. Obesity Research. 2004;12:1397–1407. - PubMed
-
- Allison DB, Pietrobelli A, Faith MS, Fontaine KR, Gropp E, et al. Genetic influences on obesity. In: Eckel RH, editor. Obesity-mechanisms and clinical management. New York: Lippincott Williams & Willkins; 2002.
-
- Atchley WR, Riska B, Kohn L, Plummet A, Rutledge JJ. A quantitative genetic analysis of brain and body size associations, their origin and ontogeny: Data from mice. Evolution. 1984;38:1165–1179. - PubMed
-
- Brockmann GA, Bevova MR. Using mouse models to dissect the genetics of obesity. Trends Genetics. 2002;18:367–376. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

