Chromosome evolution in the Thermotogales: large-scale inversions and strain diversification of CRISPR sequences
- PMID: 16547022
- PMCID: PMC1428405
- DOI: 10.1128/JB.188.7.2364-2374.2006
Chromosome evolution in the Thermotogales: large-scale inversions and strain diversification of CRISPR sequences
Abstract
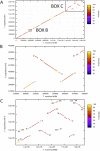
In the present study, the chromosomes of two members of the Thermotogales were compared. A whole-genome alignment of Thermotoga maritima MSB8 and Thermotoga neapolitana NS-E has revealed numerous large-scale DNA rearrangements, most of which are associated with CRISPR DNA repeats and/or tRNA genes. These DNA rearrangements do not include the putative origin of DNA replication but move within the same replichore, i.e., the same replicating half of the chromosome (delimited by the replication origin and terminus). Based on cumulative GC skew analysis, both the T. maritima and T. neapolitana lineages contain one or two major inverted DNA segments. Also, based on PCR amplification and sequence analysis of the DNA joints that are associated with the major rearrangements, the overall chromosome architecture was found to be conserved at most DNA joints for other strains of T. neapolitana. Taken together, the results from this analysis suggest that the observed chromosomal rearrangements in the Thermotogales likely occurred by successive inversions after their divergence from a common ancestor and before strain diversification. Finally, sequence analysis shows that size polymorphisms in the DNA joints associated with CRISPRs can be explained by expansion and possibly contraction of the DNA repeat and spacer unit, providing a tool for discerning the relatedness of strains from different geographic locations.
Figures




References
-
- Akimkina, T., P. Ivanov, S. Kostrov, T. Sokolova, E. Bonch-Osmolovskaya, K. Firman, C. F. Dutta, and J. A. McClellan. 1999. A highly conserved plasmid from the extreme thermophile Thermotoga maritima MC24 is a member of a family of plasmids distributed worldwide. Plasmid 42:236-240. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Bolotin, A., B. Quinquis, A. Sorokin, and S. D. Ehrlich. 2005. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551-2561. - PubMed
-
- Canchaya, C., G. Fournous, S. Chibani-Chennoufi, M. L. Dillmann, and H. Brussow. 2003. Phage as agents of lateral gene transfer. Curr. Opin. Microbiol. 6:417-424. - PubMed
-
- Chhabra, S. R., K. R. Shockley, S. B. Conners, K. L. Scott, R. D. Wolfinger, and R. M. Kelly. 2003. Carbohydrate-induced differential gene expression patterns in the hyperthermophilic bacterium Thermotoga maritima J. Biol. Chem. 278:7540-7552. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

