Exploring dynamics of protein structure determination and homology-based prediction to estimate the number of superfamilies and folds
- PMID: 16549009
- PMCID: PMC1444916
- DOI: 10.1186/1472-6807-6-6
Exploring dynamics of protein structure determination and homology-based prediction to estimate the number of superfamilies and folds
Abstract
Background: As tertiary structure is currently available only for a fraction of known protein families, it is important to assess what parts of sequence space have been structurally characterized. We consider protein domains whose structure can be predicted by sequence similarity to proteins with solved structure and address the following questions. Do these domains represent an unbiased random sample of all sequence families? Do targets solved by structural genomic initiatives (SGI) provide such a sample? What are approximate total numbers of structure-based superfamilies and folds among soluble globular domains?
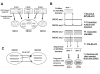
Results: To make these assessments, we combine two approaches: (i) sequence analysis and homology-based structure prediction for proteins from complete genomes; and (ii) monitoring dynamics of the assigned structure set in time, with the accumulation of experimentally solved structures. In the Clusters of Orthologous Groups (COG) database, we map the growing population of structurally characterized domain families onto the network of sequence-based connections between domains. This mapping reveals a systematic bias suggesting that target families for structure determination tend to be located in highly populated areas of sequence space. In contrast, the subset of domains whose structure is initially inferred by SGI is similar to a random sample from the whole population. To accommodate for the observed bias, we propose a new non-parametric approach to the estimation of the total numbers of structural superfamilies and folds, which does not rely on a specific model of the sampling process. Based on dynamics of robust distribution-based parameters in the growing set of structure predictions, we estimate the total numbers of superfamilies and folds among soluble globular proteins in the COG database.
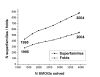
Conclusion: The set of currently solved protein structures allows for structure prediction in approximately a third of sequence-based domain families. The choice of targets for structure determination is biased towards domains with many sequence-based homologs. The growing SGI output in the future should further contribute to the reduction of this bias. The total number of structural superfamilies and folds in the COG database are estimated as approximately 4000 and approximately 1700. These numbers are respectively four and three times higher than the numbers of superfamilies and folds that can currently be assigned to COG proteins.
Figures
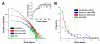


Similar articles
-
Progress of structural genomics initiatives: an analysis of solved target structures.J Mol Biol. 2005 May 20;348(5):1235-60. doi: 10.1016/j.jmb.2005.03.037. Epub 2005 Apr 2. J Mol Biol. 2005. PMID: 15854658
-
GenDiS: Genomic Distribution of protein structural domain Superfamilies.Nucleic Acids Res. 2005 Jan 1;33(Database issue):D252-5. doi: 10.1093/nar/gki087. Nucleic Acids Res. 2005. PMID: 15608190 Free PMC article.
-
Defining the fold space of membrane proteins: the CAMPS database.Proteins. 2006 Sep 1;64(4):906-22. doi: 10.1002/prot.21081. Proteins. 2006. PMID: 16802318
-
The structure of the protein universe and genome evolution.Nature. 2002 Nov 14;420(6912):218-23. doi: 10.1038/nature01256. Nature. 2002. PMID: 12432406 Review.
-
Comparing genomes in terms of protein structure: surveys of a finite parts list.FEMS Microbiol Rev. 1998 Oct;22(4):277-304. doi: 10.1111/j.1574-6976.1998.tb00371.x. FEMS Microbiol Rev. 1998. PMID: 10357579 Review.
Cited by
-
A comprehensive system for evaluation of remote sequence similarity detection.BMC Bioinformatics. 2007 Aug 28;8:314. doi: 10.1186/1471-2105-8-314. BMC Bioinformatics. 2007. PMID: 17725841 Free PMC article.
-
Small Molecule Wnt Pathway Modulators from Natural Sources: History, State of the Art and Perspectives.Cells. 2020 Mar 2;9(3):589. doi: 10.3390/cells9030589. Cells. 2020. PMID: 32131438 Free PMC article. Review.
-
Cholera- and anthrax-like toxins are among several new ADP-ribosyltransferases.PLoS Comput Biol. 2010 Dec 9;6(12):e1001029. doi: 10.1371/journal.pcbi.1001029. PLoS Comput Biol. 2010. PMID: 21170356 Free PMC article.
-
Towards a comprehensive structural coverage of completed genomes: a structural genomics viewpoint.BMC Bioinformatics. 2007 Mar 9;8:86. doi: 10.1186/1471-2105-8-86. BMC Bioinformatics. 2007. PMID: 17349043 Free PMC article.
-
Preservation of protein clefts in comparative models.BMC Struct Biol. 2008 Jan 16;8:2. doi: 10.1186/1472-6807-8-2. BMC Struct Biol. 2008. PMID: 18199319 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

