GEM System: automatic prototyping of cell-wide metabolic pathway models from genomes
- PMID: 16553966
- PMCID: PMC1435936
- DOI: 10.1186/1471-2105-7-168
GEM System: automatic prototyping of cell-wide metabolic pathway models from genomes
Abstract
Background: Successful realization of a "systems biology" approach to analyzing cells is a grand challenge for our understanding of life. However, current modeling approaches to cell simulation are labor-intensive, manual affairs, and therefore constitute a major bottleneck in the evolution of computational cell biology.
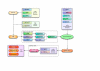
Results: We developed the Genome-based Modeling (GEM) System for the purpose of automatically prototyping simulation models of cell-wide metabolic pathways from genome sequences and other public biological information. Models generated by the GEM System include an entire Escherichia coli metabolism model comprising 968 reactions of 1195 metabolites, achieving 100% coverage when compared with the KEGG database, 92.38% with the EcoCyc database, and 95.06% with iJR904 genome-scale model.
Conclusion: The GEM System prototypes qualitative models to reduce the labor-intensive tasks required for systems biology research. Models of over 90 bacterial genomes are available at our web site.
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials