A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA
- PMID: 16585517
- PMCID: PMC1458645
- DOI: 10.1073/pnas.0509723103
A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA
Abstract
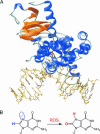
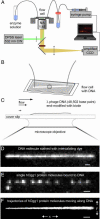
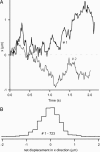
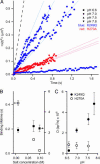
A central mystery in the function of site-specific DNA-binding proteins is the detailed mechanism for rapid location and binding of target sites in DNA. Human oxoguanine DNA glycosylase 1 (hOgg1), for example, must search out rare 8-oxoguanine lesions to prevent transversion mutations arising from oxidative stress. Here we report high-speed imaging of single hOgg1 enzyme molecules diffusing along DNA stretched by shear flow. Salt-concentration-dependent measurements reveal that such diffusion occurs as hOgg1 slides in persistent contact with DNA. At near-physiologic pH and salt concentration, hOgg1 has a subsecond DNA-binding time and slides with a diffusion constant as high as 5 x 10(6) bp(2)/s. Such a value approaches the theoretical upper limit for one-dimensional diffusion and indicates an activation barrier for sliding of only 0.5 kcal/mol (1 kcal = 4.2 kJ). This nearly barrierless Brownian sliding indicates that DNA glycosylases locate lesion bases by a massively redundant search in which the enzyme selectively binds 8-oxoguanine under kinetic control.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

