Novel hydrophobic surface binding protein, HsbA, produced by Aspergillus oryzae
- PMID: 16597938
- PMCID: PMC1449009
- DOI: 10.1128/AEM.72.4.2407-2413.2006
Novel hydrophobic surface binding protein, HsbA, produced by Aspergillus oryzae
Abstract
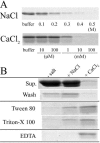
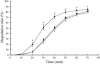
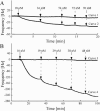
Hydrophobic surface binding protein A (HsbA) is a secreted protein (14.5 kDa) isolated from the culture broth of Aspergillus oryzae RIB40 grown in a medium containing polybutylene succinate-co-adipate (PBSA) as a sole carbon source. We purified HsbA from the culture broth and determined its N-terminal amino acid sequence. We found a DNA sequence encoding a protein whose N terminus matched that of purified HsbA in the A. ozyzae genomic sequence. We cloned the hsbA genomic DNA and cDNA from A. oryzae and constructed a recombinant A. oryzae strain highly expressing hsbA. Orthologues of HsbA were present in animal pathogenic and entomopathogenic fungi. Heterologously synthesized HsbA was purified and biochemically characterized. Although the HsbA amino acid sequence suggests that HsbA may be hydrophilic, HsbA adsorbed to hydrophobic PBSA surfaces in the presence of NaCl or CaCl(2). When HsbA was adsorbed on the hydrophobic PBSA surfaces, it promoted PBSA degradation via the CutL1 polyesterase. CutL1 interacts directly with HsbA attached to the hydrophobic QCM electrode surface. These results suggest that when HsbA is adsorbed onto the PBSA surface, it recruits CutL1, and that when CutL1 is accumulated on the PBSA surface, it stimulates PBSA degradation. We previously reported that when the A. oryzae hydrophobin RolA is bound to PBSA surfaces, it too specifically recruits CutL1. Since HsbA is not a hydrophobin, A. oryzae may use several types of proteins to recruit lytic enzymes to the surface of hydrophobic solid materials and promote their degradation.
Figures


Similar articles
-
The fungal hydrophobin RolA recruits polyesterase and laterally moves on hydrophobic surfaces.Mol Microbiol. 2005 Sep;57(6):1780-96. doi: 10.1111/j.1365-2958.2005.04803.x. Mol Microbiol. 2005. PMID: 16135240
-
Ionic interaction of positive amino acid residues of fungal hydrophobin RolA with acidic amino acid residues of cutinase CutL1.Mol Microbiol. 2015 Apr;96(1):14-27. doi: 10.1111/mmi.12915. Epub 2015 Feb 11. Mol Microbiol. 2015. PMID: 25588312
-
Analysis of the ionic interaction between the hydrophobin RodA and two cutinases of Aspergillus nidulans obtained via an Aspergillus oryzae expression system.Appl Microbiol Biotechnol. 2017 Mar;101(6):2343-2356. doi: 10.1007/s00253-016-7979-5. Epub 2016 Dec 5. Appl Microbiol Biotechnol. 2017. PMID: 27917435
-
Genomics of Aspergillus oryzae: learning from the history of Koji mold and exploration of its future.DNA Res. 2008 Aug;15(4):173-83. doi: 10.1093/dnares/dsn020. DNA Res. 2008. PMID: 18820080 Free PMC article. Review.
-
Impact of Aspergillus oryzae genomics on industrial production of metabolites.Mycopathologia. 2006 Sep;162(3):143-53. doi: 10.1007/s11046-006-0049-2. Mycopathologia. 2006. PMID: 16944282 Review.
Cited by
-
Seeking the Roles for Fungal Small-Secreted Proteins in Affecting Saprophytic Lifestyles.Front Microbiol. 2020 Mar 24;11:455. doi: 10.3389/fmicb.2020.00455. eCollection 2020. Front Microbiol. 2020. PMID: 32265881 Free PMC article. Review.
-
Partial secretome analysis of Caldariomyces fumago reveals extracellular production of the CPO co-substrate H2O2 and provides a coproduction concept for CPO and glucose oxidase.World J Microbiol Biotechnol. 2018 Jan 10;34(2):24. doi: 10.1007/s11274-017-2407-2. World J Microbiol Biotechnol. 2018. PMID: 29322262
-
Biosphere Plastic Contamination and Microbial Alternatives for a Sustainable Degradation of Plastic Waste.Microorganisms. 2025 May 28;13(6):1246. doi: 10.3390/microorganisms13061246. Microorganisms. 2025. PMID: 40572133 Free PMC article. Review.
-
Induction and Repression of Hydrolase Genes in Aspergillus oryzae.Front Microbiol. 2021 May 24;12:677603. doi: 10.3389/fmicb.2021.677603. eCollection 2021. Front Microbiol. 2021. PMID: 34108952 Free PMC article. Review.
-
Genome-wide transcriptional profiling of appressorium development by the rice blast fungus Magnaporthe oryzae.PLoS Pathog. 2012 Feb;8(2):e1002514. doi: 10.1371/journal.ppat.1002514. Epub 2012 Feb 9. PLoS Pathog. 2012. PMID: 22346750 Free PMC article.
References
-
- Ando, Y., K. Yoshikawa, T. Yoshikawa, M. Nshioka, R. Ishioka, and Y. Yakabe. 1998. Biodegradability of poly(tetramethlene scuccinate-co-tetramethylene adipate): I. Enzymatic hydrolysis. Polym. Degrad. Stab. 61:129-137.
-
- Christensen, T. 1994. Application: Aspergillus oryzae as a host for production of industrial enzymes, p. 251-260. In K. A. Powell (ed.), The genus Aspergillus. Plenum Press, New York, N.Y.
-
- De Vries, O. M. H., M. P. Fekkes, H. A. B. Wosten, and J. G. H. Wessels. 1993. Insoluble hydrophobin complexes in the walls of Schizophyllum commune and other filamentous fungi. Arch. Microbiol. 159:330-335.
-
- Fujimaki, T. 1998. Processability and properties of aliphatic polyesters, ‘BIONOLLE,’ synthesized by polycondensation reaction. Polym. Degrad. Stab. 59:209-214.
-
- Furukawa, K., Y. Hoshi, T. Maeda, T. Nakajima, and K. Abe. 2005. Aspergillus nidulans HOG pathway is activated only by two-component signaling pathway in response to osmotic stress. Mol. Microbiol. 56:1246-1261. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

