Pseudomonad cyclopentadecanone monooxygenase displaying an uncommon spectrum of Baeyer-Villiger oxidations of cyclic ketones
- PMID: 16597975
- PMCID: PMC1449013
- DOI: 10.1128/AEM.72.4.2707-2720.2006
Pseudomonad cyclopentadecanone monooxygenase displaying an uncommon spectrum of Baeyer-Villiger oxidations of cyclic ketones
Abstract
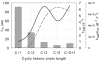
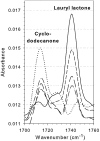
Baeyer-Villiger monooxygenases (BVMOs) are biocatalysts that offer the prospect of high chemo-, regio-, and enantioselectivity in the organic synthesis of lactones or esters from a variety of ketones. In this study, we have cloned, sequenced, and overexpressed in Escherichia coli a new BVMO, cyclopentadecanone monooxygenase (CpdB or CPDMO), originally derived from Pseudomonas sp. strain HI-70. The 601-residue primary structure of CpdB revealed only 29% to 50% sequence identity to those of known BVMOs. A new sequence motif, characterized by a cluster of charged residues, was identified in a subset of BVMO sequences that contain an N-terminal extension of approximately 60 to 147 amino acids. The 64-kDa CPDMO enzyme was purified to apparent homogeneity, providing a specific activity of 3.94 micromol/min/mg protein and a 20% yield. CPDMO is monomeric and NADPH dependent and contains approximately 1 mol flavin adenine dinucleotide per mole of protein. A deletion mutant suggested the importance of the N-terminal 54 amino acids to CPDMO activity. In addition, a Ser261Ala substitution in a Rossmann fold motif resulted in an improved stability and increased affinity of the enzyme towards NADPH compared to the wild-type enzyme (K(m) = 8 microM versus K(m) = 24 microM). Substrate profiling indicated that CPDMO is unusual among known BVMOs in being able to accommodate and oxidize both large and small ring substrates that include C(11) to C(15) ketones, methyl-substituted C(5) and C(6) ketones, and bicyclic ketones, such as decalone and beta-tetralone. CPDMO has the highest affinity (K(m) = 5.8 microM) and the highest catalytic efficiency (k(cat)/K(m) ratio of 7.2 x 10(5) M(-1) s(-1)) toward cyclopentadecanone, hence the Cpd designation. A number of whole-cell biotransformations were carried out, and as a result, CPDMO was found to have an excellent enantioselectivity (E > 200) as well as 99% S-selectivity toward 2-methylcyclohexanone for the production of 7-methyl-2-oxepanone, a potentially valuable chiral building block. Although showing a modest selectivity (E = 5.8), macrolactone formation of 15-hexadecanolide from the kinetic resolution of 2-methylcyclopentadecanone using CPDMO was also demonstrated.
Figures
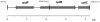


References
-
- Alphand, V., G. Carrea, R. Wohlgemuth, R. Furstoss, and J. M. Woodley. 2003. Towards large-scale synthetic applications of Baeyer-Villiger monooxygenases. Trends Biotechnol. 21:318-323. - PubMed
-
- Ayala, M., and E. Torres. 2004. Enzymatic activation of alkanes: constraints and prospective. Appl. Catal. A 272:1-13.
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72:248-254. - PubMed
-
- Brzostowicz, P. C., M. S. Blasko, and P. E. Rouviere. 2002. Identification of two gene clusters involved in cyclohexanone oxidation in Brevibacterium epidermidis strain HCU. Appl. Microbiol. Biotechnol. 58:781-789. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

