Bacterial display using circularly permuted outer membrane protein OmpX yields high affinity peptide ligands
- PMID: 16600968
- PMCID: PMC2242469
- DOI: 10.1110/ps.051897806
Bacterial display using circularly permuted outer membrane protein OmpX yields high affinity peptide ligands
Abstract
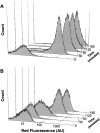
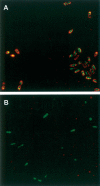
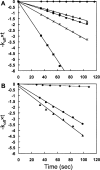
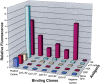
A bacterial display methodology was developed for N- and C-terminal display and demonstrated to enable rapid screening of very large peptide libraries with high precision and efficiency. To overcome limitations of insertional fusion display libraries, a new scaffold was developed through circular permutation of the Escherichia coli outer membrane protein OmpX that presents both N and C termini on the external cell surface. Circularly permuted OmpX (CPX) display was directly compared to insertional fusion display by screening comparable peptide libraries in each format using magnetic and fluorescence activated cell sorting. CPX display enabled in situ measurement of dissociation rate constants with improved accuracy and, consequently, improved affinity discrimination during screening and ranking of isolated clones. Using streptavidin as a model target, bacterial display yielded the well-characterized HP(Q)/(M) motif obtained previously using several alternative peptide display systems, as well as three additional motifs (L(I)/(V) CQNVCY, CGWMY(F)/(Y)xEC, ERCWYVMHWPCNA). Using CPX display, a very high affinity streptavidin-binding peptide was isolated having a dissociation rate constant k(off) = 0.002sec(-1) even after grafting to the C terminus of an unrelated protein. Comparison of individual clones obtained from insertional fusion and terminal fusion libraries suggests that the N-terminal display yields sequences with greater diversity, affinity, and modularity. CPX bacterial display thus provides a highly effective method for screening peptide libraries to rapidly generate ligands with high affinity and specificity.
Figures


Similar articles
-
Substrate specificity of the Escherichia coli outer membrane protease OmpP.J Bacteriol. 2007 Jan;189(2):522-30. doi: 10.1128/JB.01493-06. Epub 2006 Nov 3. J Bacteriol. 2007. PMID: 17085556 Free PMC article.
-
Rapid isolation of high-affinity protein binding peptides using bacterial display.Protein Eng Des Sel. 2004 Oct;17(10):731-9. doi: 10.1093/protein/gzh084. Epub 2004 Nov 5. Protein Eng Des Sel. 2004. PMID: 15531628
-
Directed evolution of a biterminal bacterial display scaffold enhances the display of diverse peptides.Protein Eng Des Sel. 2008 Jul;21(7):435-42. doi: 10.1093/protein/gzn020. Epub 2008 May 13. Protein Eng Des Sel. 2008. PMID: 18480093 Free PMC article.
-
Decorating microbes: surface display of proteins on Escherichia coli.Trends Biotechnol. 2011 Feb;29(2):79-86. doi: 10.1016/j.tibtech.2010.11.003. Epub 2010 Dec 9. Trends Biotechnol. 2011. PMID: 21146237 Review.
-
Insertional gene fusion technology.FEBS Lett. 1999 Aug 20;457(1):1-4. doi: 10.1016/s0014-5793(99)00991-6. FEBS Lett. 1999. PMID: 10486551 Review.
Cited by
-
The next generation of biopanning: next gen sequencing improves analysis of bacterial display libraries.BMC Biotechnol. 2019 Dec 21;19(1):100. doi: 10.1186/s12896-019-0577-8. BMC Biotechnol. 2019. PMID: 31864334 Free PMC article.
-
Artificial surface labelling of Escherichia coli with StrepTagII antigen to study how monoclonal antibodies drive complement-mediated killing.Sci Rep. 2023 Nov 1;13(1):18836. doi: 10.1038/s41598-023-46026-x. Sci Rep. 2023. PMID: 37914798 Free PMC article.
-
Screening of peptide libraries against protective antigen of Bacillus anthracis in a disposable microfluidic cartridge.PLoS One. 2011;6(11):e26925. doi: 10.1371/journal.pone.0026925. Epub 2011 Nov 28. PLoS One. 2011. PMID: 22140433 Free PMC article.
-
Surface display of recombinant proteins on Escherichia coli by BclA exosporium of Bacillus anthracis.Microb Cell Fact. 2013 Sep 22;12:81. doi: 10.1186/1475-2859-12-81. Microb Cell Fact. 2013. PMID: 24053632 Free PMC article.
-
Exploiting bacterial peptide display technology to engineer biomaterials for neural stem cell culture.Biomaterials. 2011 Feb;32(6):1484-94. doi: 10.1016/j.biomaterials.2010.10.032. Epub 2010 Dec 3. Biomaterials. 2011. PMID: 21129772 Free PMC article.
References
-
- Becker S., Theile S., Heppeler N., Michalczyk A., Wentzel A., Wilhelm S., Jaeger K.E., Kolmar H. 2005. A generic system for the Escherichia coli cell-surface display of lipolytic enzymes FEBS Lett. 579: 1177–1182. - PubMed
-
- Bessette P.H. and Daugherty P.S. 2004. Flow cytometric screening of cDNA expression libraries for fluorescent proteins Biotechnol. Prog. 20: 963–967. - PubMed
-
- Bessette P.H., Rice J.J., Daugherty P.S. 2004. Rapid isolation of high-affinity protein binding peptides using bacterial display Protein Eng. Des. Sel. 17: 731–739. - PubMed
-
- Boder E.T. and Wittrup K.D. 1997. Yeast surface display for screening combinatorial polypeptide libraries Nat. Biotechnol. 15: 553–557. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

