Profiling alternatively spliced mRNA isoforms for prostate cancer classification
- PMID: 16608523
- PMCID: PMC1458362
- DOI: 10.1186/1471-2105-7-202
Profiling alternatively spliced mRNA isoforms for prostate cancer classification
Abstract
Background: Prostate cancer is one of the leading causes of cancer illness and death among men in the United States and world wide. There is an urgent need to discover good biomarkers for early clinical diagnosis and treatment. Previously, we developed an exon-junction microarray-based assay and profiled 1532 mRNA splice isoforms from 364 potential prostate cancer related genes in 38 prostate tissues. Here, we investigate the advantage of using splice isoforms, which couple transcriptional and splicing regulation, for cancer classification.
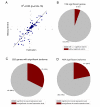
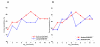
Results: As many as 464 splice isoforms from more than 200 genes are differentially regulated in tumors at a false discovery rate (FDR) of 0.05. Remarkably, about 30% of genes have isoforms that are called significant but do not exhibit differential expression at the overall mRNA level. A support vector machine (SVM) classifier trained on 128 signature isoforms can correctly predict 92% of the cases, which outperforms the classifier using overall mRNA abundance by about 5%. It is also observed that the classification performance can be improved using multivariate variable selection methods, which take correlation among variables into account.
Conclusion: These results demonstrate that profiling of splice isoforms is able to provide unique and important information which cannot be detected by conventional microarrays.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

