Interactions of mRNAs and gRNAs involved in trypanosome mitochondrial RNA editing: structure probing of a gRNA bound to its cognate mRNA
- PMID: 16618968
- PMCID: PMC1464861
- DOI: 10.1261/rna.3406
Interactions of mRNAs and gRNAs involved in trypanosome mitochondrial RNA editing: structure probing of a gRNA bound to its cognate mRNA
Abstract
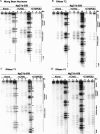
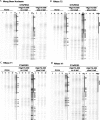
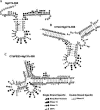
Expression of mitochondrial genes in Trypanosoma brucei requires RNA editing of its mRNA transcripts. During editing, uridylates are precisely inserted and deleted as directed by the gRNA template to create the protein open reading frame. This process involves the bimolecular interaction of the gRNA with its cognate pre-edited mRNA and the assembly of a protein complex with the enzymatic machinery required. While a considerable amount of work has been done identifying the protein components of the editing complex, very little is known about how a functional editosome is assembled. In addition, the importance of RNA structure in establishing a functional editing complex is poorly understood. Work in our lab suggests that different mRNA/gRNA pairs can form similar secondary structures suggesting that a common core architecture may be important for editosome recognition and function. Using solution structure probing, we have investigated the structure of the initiating gRNA, gCYb-558, in the mRNA/gRNA complex with pre-edited apocytochrome b mRNA. Our data indicate that the stem-loop formed by the guiding region of the gRNA alone is maintained in its interaction with the pre-edited message. In addition, our data suggest that a gRNA stem-loop structure is maintained through the first few editing events by the use of alternative base-pairing with the U-tail.
Figures
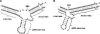



References
-
- Aphasizhev R., Simpson L. Isolation and characterization of a U-specific 3′-5′-exonuclease from mitochondria of Leishmania tarentolae . J. Biol. Chem. 2001;276:21280–21284. - PubMed
-
- Aphasizhev R., Sbicego S., Peris M., Jang S.H., Aphasizheva I., Simpson A.M., Rivlin A., Simpson L. Trypanosome mitochondrial 3′ terminal uridylyl transferase (TUTase): The key enzyme in U-insertion/deletion RNA editing. Cell. 2002;108:637–648. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
