Residues in two homology blocks on the amino side of the tRNase Z His domain contribute unexpectedly to pre-tRNA 3' end processing
- PMID: 16618969
- PMCID: PMC1464858
- DOI: 10.1261/rna.4206
Residues in two homology blocks on the amino side of the tRNase Z His domain contribute unexpectedly to pre-tRNA 3' end processing
Abstract
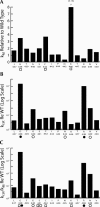
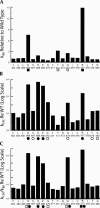
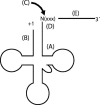
tRNase Z, which can endonucleolytically remove pre-tRNA 3'-end trailers, possesses the signature His domain (HxHxDH; Motif II) of the beta-lactamase family of metal-dependent hydrolases. Motif II combines with Motifs III-V on its carboxy side to coordinate two divalent metal ions, constituting the catalytic core. The PxKxRN loop and Motif I on the amino side of Motif II have been suggested to modulate tRNase Z activity, including the anti-determinant effect of CCA in mature tRNA. Ala walks through these two homology blocks reveal residues in which the substitutions unexpectedly reduce catalytic efficiency. While substitutions in Motif II can drastically affect k(cat) without affecting k(M), five- to 15-fold increases in k(M) are observed with substitutions in several conserved residues in the PxKxRN loop and Motif I. These increases in k(M) suggest a model for substrate binding. Expressed tRNase Z processes mature tRNA with CCA at the 3' end approximately 80 times less efficiently than a pre-tRNA possessing natural sequence of the 3'-end trailer, due to reduced k(cat) with no effect on k(M), showing the CCA anti-determinant to be a characteristic of this enzyme.
Figures





Similar articles
-
Identification and sequence analysis of metazoan tRNA 3'-end processing enzymes tRNase Zs.PLoS One. 2012;7(9):e44264. doi: 10.1371/journal.pone.0044264. Epub 2012 Sep 4. PLoS One. 2012. PMID: 22962606 Free PMC article.
-
tRNase Z catalysis and conserved residues on the carboxy side of the His cluster.Biochemistry. 2007 Aug 21;46(33):9380-7. doi: 10.1021/bi700578v. Epub 2007 Jul 27. Biochemistry. 2007. PMID: 17655328 Free PMC article.
-
Residues in the conserved His domain of fruit fly tRNase Z that function in catalysis are not involved in substrate recognition or binding.J Mol Biol. 2005 Jul 8;350(2):189-99. doi: 10.1016/j.jmb.2005.04.073. J Mol Biol. 2005. PMID: 15935379
-
tRNase Z: the end is not in sight.Cell Mol Life Sci. 2007 Sep;64(18):2404-12. doi: 10.1007/s00018-007-7160-5. Cell Mol Life Sci. 2007. PMID: 17599240 Free PMC article. Review.
-
The making of tRNAs and more - RNase P and tRNase Z.Prog Mol Biol Transl Sci. 2009;85:319-68. doi: 10.1016/S0079-6603(08)00808-8. Prog Mol Biol Transl Sci. 2009. PMID: 19215776 Review.
Cited by
-
Identification and sequence analysis of metazoan tRNA 3'-end processing enzymes tRNase Zs.PLoS One. 2012;7(9):e44264. doi: 10.1371/journal.pone.0044264. Epub 2012 Sep 4. PLoS One. 2012. PMID: 22962606 Free PMC article.
-
Catalytic properties of RNase BN/RNase Z from Escherichia coli: RNase BN is both an exo- and endoribonuclease.J Biol Chem. 2009 Jun 5;284(23):15425-31. doi: 10.1074/jbc.M109.005462. Epub 2009 Apr 14. J Biol Chem. 2009. PMID: 19366704 Free PMC article.
-
A survey of green plant tRNA 3'-end processing enzyme tRNase Zs, homologs of the candidate prostate cancer susceptibility protein ELAC2.BMC Evol Biol. 2011 Jul 23;11:219. doi: 10.1186/1471-2148-11-219. BMC Evol Biol. 2011. PMID: 21781332 Free PMC article.
-
tRNase Z catalysis and conserved residues on the carboxy side of the His cluster.Biochemistry. 2007 Aug 21;46(33):9380-7. doi: 10.1021/bi700578v. Epub 2007 Jul 27. Biochemistry. 2007. PMID: 17655328 Free PMC article.
-
Identification and analysis of candidate fungal tRNA 3'-end processing endonucleases tRNase Zs, homologs of the putative prostate cancer susceptibility protein ELAC2.BMC Evol Biol. 2010 Sep 6;10:272. doi: 10.1186/1471-2148-10-272. BMC Evol Biol. 2010. PMID: 20819227 Free PMC article.
References
-
- Aebi M., Kirchner G., Chen J.Y., Vijayraghavan U., Jacobson A., Martin N.C., Abelson J. Isolation of a temperature-sensitive mutant with an altered tRNA nucleotidyltransferase and cloning of the gene encoding tRNA nucleotidyltransferase in the yeast Saccharomyces cerevisiae . J. Biol. Chem. 1990;265:16216–16220. - PubMed
-
- Baillat D., Hakimi M.A., Naar A.M., Shilatifard A., Cooch N., Shiekhattar R. Integrator, a multiprotein mediator of small nuclear RNA processing, associates with the C-terminal repeat of RNA polymerase II. Cell. 2005;123:265–276. - PubMed
-
- Castaño J.G., Tobian J.A., Zasloff M. Purification and characterization of an endonuclease from Xenopus laevis ovaries which accurately processes the 3′ terminus of human pre-tRNA-Met(i) (3′ pre-tRNase) J. Biol. Chem. 1985;260:9002–9008. - PubMed
-
- Chen J.Y., Martin N.C. Biosynthesis of tRNA in yeast mitochondria. An endonuclease is responsible for the 3′-processing of tRNA precursors. J. Biol. Chem. 1988;263:13677–13682. - PubMed
-
- de la Sierra-Gallay I.L., Pellegrini O., Condon C. Structural basis for substrate binding, cleavage and allostery in the tRNA maturase RNase Z. Nature. 2005;433:657–661. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
