Arrested yeast splicing complexes indicate stepwise snRNP recruitment during in vivo spliceosome assembly
- PMID: 16618970
- PMCID: PMC1464846
- DOI: 10.1261/rna.50506
Arrested yeast splicing complexes indicate stepwise snRNP recruitment during in vivo spliceosome assembly
Abstract
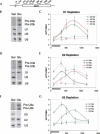
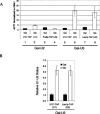
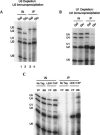
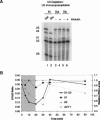
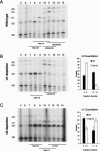
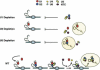
Pre-mRNA splicing is catalyzed by the spliceosome, a macromolecular machine dedicated to intron removal and exon ligation. Despite an abundance of in vitro information and a small number of in vivo studies, the pathway of yeast (Saccharomyces cerevisiae) in vivo spliceosome assembly remains uncertain. To address this situation, we combined in vivo depletions of U1, U2, or U5 snRNAs with chromatin immunoprecipitation (ChIP) analysis of other splicing snRNPs along an intron-containing gene. The data indicate that snRNP recruitment to nascent pre-mRNA predominantly proceeds via the canonical three-step assembly pathway: first U1, then U2, and finally the U4/U6*U5 tri-snRNP. Tandem affinity purification (TAP) using a U2 snRNP-tagged protein allowed the characterization of in vivo assembled higher-order splicing complexes. Consistent with an independent snRNP assembly pathway, we observed high levels of U1-U2 prespliceosomes under U5-depletion conditions, and we observed significant levels of a U2/U5/U6/Prp19-complex mature splicing complex under wild-type conditions. These complexes have implications for the steady-state distribution of snRNPs within nuclei and also reinforce the stepwise recruitment of U1, U2, and the tri-snRNP during in vivo spliceosome assembly.
Figures
Similar articles
-
Prespliceosome structure provides insights into spliceosome assembly and regulation.Nature. 2018 Jul;559(7714):419-422. doi: 10.1038/s41586-018-0323-8. Epub 2018 Jul 11. Nature. 2018. PMID: 29995849 Free PMC article.
-
A novel yeast U2 snRNP protein, Snu17p, is required for the first catalytic step of splicing and for progression of spliceosome assembly.Mol Cell Biol. 2001 May;21(9):3037-46. doi: 10.1128/MCB.21.9.3037-3046.2001. Mol Cell Biol. 2001. PMID: 11287609 Free PMC article.
-
The splicing factor Prp17 interacts with the U2, U5 and U6 snRNPs and associates with the spliceosome pre- and post-catalysis.Biochem J. 2008 Dec 15;416(3):365-74. doi: 10.1042/BJ20081195. Biochem J. 2008. PMID: 18691155
-
CryoEM structures of two spliceosomal complexes: starter and dessert at the spliceosome feast.Curr Opin Struct Biol. 2016 Feb;36:48-57. doi: 10.1016/j.sbi.2015.12.005. Epub 2016 Jan 21. Curr Opin Struct Biol. 2016. PMID: 26803803 Free PMC article. Review.
-
Branch site recognition by the spliceosome.RNA. 2024 Oct 16;30(11):1397-1407. doi: 10.1261/rna.080198.124. RNA. 2024. PMID: 39187383 Free PMC article. Review.
Cited by
-
Tailoring of membrane proteins by alternative splicing of pre-mRNA.Biochemistry. 2012 Jul 17;51(28):5541-56. doi: 10.1021/bi3007065. Epub 2012 Jun 29. Biochemistry. 2012. PMID: 22708632 Free PMC article. Review.
-
Single molecule analysis reveals reversible and irreversible steps during spliceosome activation.Elife. 2016 May 31;5:e14166. doi: 10.7554/eLife.14166. Elife. 2016. PMID: 27244240 Free PMC article.
-
Nascent RNA and the Coordination of Splicing with Transcription.Cold Spring Harb Perspect Biol. 2019 Aug 1;11(8):a032227. doi: 10.1101/cshperspect.a032227. Cold Spring Harb Perspect Biol. 2019. PMID: 31371351 Free PMC article. Review.
-
The differential interaction of snRNPs with pre-mRNA reveals splicing kinetics in living cells.J Cell Biol. 2010 Oct 4;191(1):75-86. doi: 10.1083/jcb.201004030. J Cell Biol. 2010. PMID: 20921136 Free PMC article.
-
The abundance of the spliceosomal snRNPs is not limiting the splicing of U12-type introns.RNA. 2006 Oct;12(10):1883-92. doi: 10.1261/rna.213906. Epub 2006 Sep 6. RNA. 2006. PMID: 16957280 Free PMC article.
References
-
- Burge C.B., Tuschl T., Sharp P.A. Splicing of precursors to mRNAs by the spliceosome. In: Gesteland R.R., et al., editors. The RNA world. 2d ed. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY: 1999. pp. 525–560.
-
- Das R., Zhou Z., Reed R. Functional association of U2 snRNP with the ATP-independent spliceosomal complex E. Mol. Cell. 2000;5:779–787. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
