Role of sigmaD in regulating genes and signals during Myxococcus xanthus development
- PMID: 16621817
- PMCID: PMC1447441
- DOI: 10.1128/JB.188.9.3246-3256.2006
Role of sigmaD in regulating genes and signals during Myxococcus xanthus development
Abstract
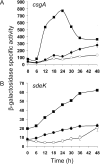
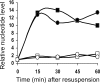
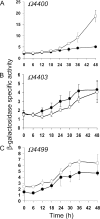
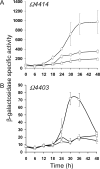
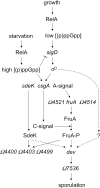
Starvation-induced development of Myxococcus xanthus is an excellent model for biofilm formation because it involves cell-cell signaling to coordinate formation of multicellular mounds, gene expression, and cellular differentiation into spores. The role of sigma(D), an alternative sigma factor important for viability in stationary phase and for stress responses, was investigated during development by measuring signal production, gene expression, and sporulation of a sigD null mutant alone and upon codevelopment with wild-type cells or signaling mutants. The sigD mutant responded to starvation by inducing (p)ppGpp synthesis normally but was impaired for production of A-signal, an early cell density signal, and for production of the morphogenetic C-signal. Induction of early developmental genes was greatly reduced, and expression of those that depend on A-signal was not restored by codevelopment with wild-type cells, indicating that sigma(D) is needed for cellular responses to A-signal. Despite these early developmental defects, the sigD mutant responded to C-signal supplied by codeveloping wild-type cells by inducing a subset of late developmental genes. sigma(D) RNA polymerase is dispensable for transcription of this subset, but a distinct regulatory class, which includes genes essential for sporulation, requires sigma(D) RNA polymerase or a gene under its control, cell autonomously. The level of sigD transcript in a relA mutant during growth is much lower than in wild-type cells, suggesting that (p)ppGpp positively regulates sigD transcription in growing cells. The sigD transcript level drops in wild-type cells after 20 min of starvation and remains low after 40 min but rises in a relA mutant after 40 min, suggesting that (p)ppGpp negatively regulates sigD transcription early in development. We conclude that sigma(D) synthesized during growth occupies a position near the top of a regulatory hierarchy governing M. xanthus development, analogous to sigma factors that control biofilm formation of other bacteria.
Figures



Similar articles
-
A new sigma factor, SigD, essential for stationary phase is also required for multicellular differentiation in Myxococcus xanthus.Genes Cells. 1998 Jun;3(6):371-85. doi: 10.1046/j.1365-2443.1998.00197.x. Genes Cells. 1998. PMID: 9734783
-
The dev Operon Regulates the Timing of Sporulation during Myxococcus xanthus Development.J Bacteriol. 2017 Apr 25;199(10):e00788-16. doi: 10.1128/JB.00788-16. Print 2017 May 15. J Bacteriol. 2017. PMID: 28264995 Free PMC article.
-
Short-range C-signaling restricts cheating behavior during Myxococcus xanthus development.mBio. 2024 Nov 13;15(11):e0244024. doi: 10.1128/mbio.02440-24. Epub 2024 Oct 18. mBio. 2024. PMID: 39422488 Free PMC article.
-
Highly Signal-Responsive Gene Regulatory Network Governing Myxococcus Development.Trends Genet. 2017 Jan;33(1):3-15. doi: 10.1016/j.tig.2016.10.006. Epub 2016 Dec 2. Trends Genet. 2017. PMID: 27916428 Free PMC article. Review.
-
Cell-cell interactions that direct fruiting body development in Myxococcus xanthus.Curr Opin Genet Dev. 1991 Oct;1(3):363-9. doi: 10.1016/s0959-437x(05)80301-6. Curr Opin Genet Dev. 1991. PMID: 1840894 Review.
Cited by
-
Regulated proteolysis in bacterial development.FEMS Microbiol Rev. 2014 May;38(3):493-522. doi: 10.1111/1574-6976.12050. Epub 2013 Dec 19. FEMS Microbiol Rev. 2014. PMID: 24354618 Free PMC article. Review.
-
Proteomic analysis of stationary phase in the marine bacterium "Candidatus Pelagibacter ubique".Appl Environ Microbiol. 2008 Jul;74(13):4091-100. doi: 10.1128/AEM.00599-08. Epub 2008 May 9. Appl Environ Microbiol. 2008. PMID: 18469119 Free PMC article.
-
Regulation of dev, an operon that includes genes essential for Myxococcus xanthus development and CRISPR-associated genes and repeats.J Bacteriol. 2007 May;189(10):3738-50. doi: 10.1128/JB.00187-07. Epub 2007 Mar 16. J Bacteriol. 2007. PMID: 17369305 Free PMC article.
-
Identification and characterization of a putative arginine kinase homolog from Myxococcus xanthus required for fruiting body formation and cell differentiation.J Bacteriol. 2012 May;194(10):2668-76. doi: 10.1128/JB.06435-11. Epub 2012 Mar 2. J Bacteriol. 2012. PMID: 22389486 Free PMC article.
-
Mutations of the act promoter in Myxococcus xanthus.J Bacteriol. 2007 Mar;189(5):1836-44. doi: 10.1128/JB.01618-06. Epub 2006 Dec 22. J Bacteriol. 2007. PMID: 17189369 Free PMC article.
References
-
- Aimes, B., and B. Dubin. 1960. The role of polyamines in the neutralization of bacteriophage deoxyribonucleic acid. J. Biol. Chem. 235:769-775. - PubMed
-
- Biran, D., and L. Kroos. 1997. In vitro transcription of Myxococcus xanthus genes with RNA polymerase containing σA, the major sigma factor in growing cells. Mol. Microbiol. 25:463-472. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

