DEMETER and REPRESSOR OF SILENCING 1 encode 5-methylcytosine DNA glycosylases
- PMID: 16624880
- PMCID: PMC1458983
- DOI: 10.1073/pnas.0601109103
DEMETER and REPRESSOR OF SILENCING 1 encode 5-methylcytosine DNA glycosylases
Abstract
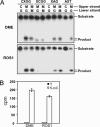
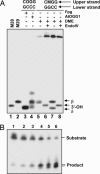
Cytosine methylation is an epigenetic mark that promotes gene silencing and plays important roles in development and genome defense against transposons. Methylation patterns are established and maintained by DNA methyltransferases that catalyze transfer of a methyl group from S-adenosyl-L-methionine to cytosine bases in DNA. Erasure of cytosine methylation occurs during development, but the enzymatic basis of active demethylation remains controversial. In Arabidopsis thaliana, DEMETER (DME) activates the maternal expression of two imprinted genes silenced by methylation, and REPRESSOR OF SILENCING 1 (ROS1) is required for release of transcriptional silencing of a hypermethylated transgene. DME and ROS1 encode two closely related DNA glycosylase domain proteins, but it is unknown whether they participate directly in a DNA demethylation process or counteract silencing through an indirect effect on chromatin structure. Here we show that DME and ROS1 catalyze the release of 5-methylcytosine (5-meC) from DNA by a glycosylase/lyase mechanism. Both enzymes also remove thymine, but not uracil, mismatched to guanine. DME and ROS1 show a preference for 5-meC over thymine in the symmetric dinucleotide CpG context, where most plant DNA methylation occurs. Nevertheless, they also have significant activity on both substrates at CpApG and asymmetric sequences, which are additional methylation targets in plant genomes. These findings suggest that a function of ROS1 and DME is to initiate erasure of 5-meC through a base excision repair process and provide strong biochemical evidence for the existence of an active DNA demethylation pathway in plants.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

