The FUSE/FBP/FIR/TFIIH system is a molecular machine programming a pulse of c-myc expression
- PMID: 16628215
- PMCID: PMC1462968
- DOI: 10.1038/sj.emboj.7601101
The FUSE/FBP/FIR/TFIIH system is a molecular machine programming a pulse of c-myc expression
Abstract
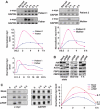
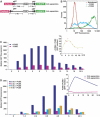
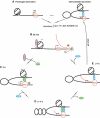
FarUpStream Element (FUSE) Binding Protein (FBP) binds the human c-myc FUSE in vitro only in single-stranded or supercoiled DNA. Because transcriptionally generated torsion melts FUSE in vitro even in linear DNA, and FBP/FBP Interacting Repressor (FIR) regulates transcription through TFIIH, these components have been speculated to be the mechanosensor (FUSE) and effectors (FBP/FIR) of a real-time mechanism controlling c-myc transcription. To ascertain whether the FUSE/FBP/FIR system operates according to this hypothesis in vivo, the flux of activators, repressors and chromatin remodeling complexes on the c-myc promoter was monitored throughout the serum-induced pulse of transcription. After transcription was switched on by conventional factors and chromatin regulators, FBP and FIR were recruited and established a dynamically remodeled loop with TFIIH at the P2 promoter. In XPB cells carrying mutant TFIIH, loop formation failed and the serum response was abnormal; RNAi depletion of FIR similarly disabled c-myc regulation. Engineering FUSE into episomal vectors predictably re-programmed metallothionein-promoter-driven reporter expression. The in vitro recruitment of FBP and FIR to dynamically stressed c-myc DNA paralleled the in vivo process.
Figures
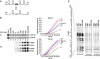
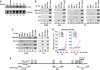

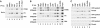
References
-
- Bazar L, Meighen D, Harris V, Duncan R, Levens D, Avigan M (1995) Targeted melting and binding of a DNA regulatory element by a transactivator of c-myc. J Biol Chem 270: 8241–8248 - PubMed
-
- Bentley DL, Groudine M (1986) A block to elongation is largely responsible for decreased transcription of c-myc in differentiated HL60 cells. Nature 321: 702–706 - PubMed
-
- Carey M, Smale ST (2000) Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY
-
- Cawley S, Bekiranov S, Ng HH, Kapranov P, Sekinger EA, Kampa D, Piccolboni A, Sementchenko V, Cheng J, Williams AJ, Wheeler R, Wong B, Drenkow J, Yamanaka M, Patel S, Brubaker S, Tammana H, Helt G, Struhl K, Gingeras TR (2004) Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116: 499–509 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

