Analysis of HIV-1 CRF_01 A/E protease inhibitor resistance: structural determinants for maintaining sensitivity and developing resistance to atazanavir
- PMID: 16634628
- PMCID: PMC2518317
- DOI: 10.1021/bi051886s
Analysis of HIV-1 CRF_01 A/E protease inhibitor resistance: structural determinants for maintaining sensitivity and developing resistance to atazanavir
Abstract
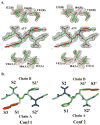
A series of HIV-1 protease mutants has been designed in an effort to analyze the contribution to drug resistance provided by natural polymorphisms as well as therapy-selective (active and non-active site) mutations in the HIV-1 CRF_01 A/E (AE) protease when compared to that of the subtype B (B) protease. Kinetic analysis of these variants using chromogenic substrates showed differences in substrate specificity between pretherapy B and AE proteases. Inhibition analysis with ritonavir, indinavir, nelfinavir, amprenavir, saquinavir, lopinavir, and atazanavir revealed that the natural polymorphisms found in A/E can influence inhibitor resistance. It was also apparent that a high level of resistance in the A/E protease, as with B protease, is due to it aquiring a combination of active site and non-active site mutations. Structural analysis of atazanavir bound to a pretherapy B protease showed that the ability of atazanavir to maintain its binding affinity for variants containing some resistance mutations is due to its unique interactions with flap residues. This structure also explains why the I50L and I84V mutations are important in decreasing the binding affinity of atazanavir.
Figures





References
-
- Velazquez-Campoy A, Vega S, Freire E. Amplification of the effects of drug resistance mutations by background polymorphisms in HIV-1 protease from African subtypes. Biochemistry. 2002;41:8613–9. - PubMed
-
- Pieniazek D, Rayfield M, Hu DJ, Nkengasong J, Wiktor SZ, Downing R, Biryahwaho B, Mastro T, Tanuri A, Soriano V, Lal R, Dondero T. Protease sequences from HIV-1 group M subtypes A-H reveal distinct amino acid mutation patterns associated with protease resistance in protease inhibitor-naive individuals worldwide. HIV Variant Working Group. Aids. 2000;14:1489–95. - PubMed
-
- Vergne L, Peeters M, Mpoudi-Ngole E, Bourgeois A, Liegeois F, Toure-Kane C, Mboup S, Mulanga-Kabeya C, Saman E, Jourdan J, Reynes J, Delaporte E. Genetic diversity of protease and reverse transcriptase sequences in non-subtype-B human immunodeficiency virus type 1 strains: evidence of many minor drug resistance mutations in treatment-naive patients. J Clin Microbiol. 2000;38:3919–25. - PMC - PubMed
-
- Cornelissen M, van den Burg R, Zorgdrager F, Lukashov V, Goudsmit J. pol gene diversity of five human immunodeficiency virus type 1 subtypes: evidence for naturally occurring mutations that contribute to drug resistance, limited recombination patterns, and common ancestry for subtypes B and D. J Virol. 1997;71:6348–58. - PMC - PubMed
-
- Grossman Z, Vardinon N, Chemtob D, Alkan ML, Bentwich Z, Burke M, Gottesman G, Istomin V, Levi I, Maayan S, Shahar E, Schapiro JM. Genotypic variation of HIV-1 reverse transcriptase and protease: comparative analysis of clade C and clade B. Aids. 2001;15:1453–60. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Chemical Information

