Global transcriptional profiling of the toxic dinoflagellate Alexandrium fundyense using Massively Parallel Signature Sequencing
- PMID: 16638123
- PMCID: PMC1473201
- DOI: 10.1186/1471-2164-7-88
Global transcriptional profiling of the toxic dinoflagellate Alexandrium fundyense using Massively Parallel Signature Sequencing
Abstract
Background: Dinoflagellates are one of the most important classes of marine and freshwater algae, notable both for their functional diversity and ecological significance. They occur naturally as free-living cells, as endosymbionts of marine invertebrates and are well known for their involvement in "red tides". Dinoflagellates are also notable for their unusual genome content and structure, which suggests that the organization and regulation of dinoflagellate genes may be very different from that of most eukaryotes. To investigate the content and regulation of the dinoflagellate genome, we performed a global analysis of the transcriptome of the toxic dinoflagellate Alexandrium fundyense under nitrate- and phosphate-limited conditions using Massively Parallel Signature Sequencing (MPSS).
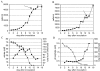
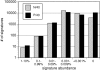
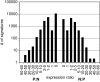
Results: Data from the two MPSS libraries showed that the number of unique signatures found in A. fundyense cells is similar to that of humans and Arabidopsis thaliana, two eukaryotes that have been extensively analyzed using this method. The general distribution, abundance and expression patterns of the A. fundyense signatures were also quite similar to other eukaryotes, and at least 10% of the A. fundyense signatures were differentially expressed between the two conditions. RACE amplification and sequencing of a subset of signatures showed that multiple signatures arose from sequence variants of a single gene. Single signatures also mapped to different sequence variants of the same gene.
Conclusion: The MPSS data presented here provide a quantitative view of the transcriptome and its regulation in these unusual single-celled eukaryotes. The observed signature abundance and distribution in Alexandrium is similar to that of other eukaryotes that have been analyzed using MPSS. Results of signature mapping via RACE indicate that many signatures result from sequence variants of individual genes. These data add to the growing body of evidence for widespread gene duplication in dinoflagellates, which would contribute to the transcriptional complexity of these organisms. The MPSS data also demonstrate that a significant number of dinoflagellate mRNAs are transcriptionally regulated, indicating that dinoflagellates commonly employ transcriptional gene regulation along with the post-transcriptional regulation that has been well documented in these organisms.
Figures

Similar articles
-
Transcriptome profiling of a toxic dinoflagellate reveals a gene-rich protist and a potential impact on gene expression due to bacterial presence.PLoS One. 2010 Mar 12;5(3):e9688. doi: 10.1371/journal.pone.0009688. PLoS One. 2010. PMID: 20300646 Free PMC article.
-
Analysis of the transcriptional complexity of Arabidopsis thaliana by massively parallel signature sequencing.Nat Biotechnol. 2004 Aug;22(8):1006-11. doi: 10.1038/nbt992. Epub 2004 Jul 11. Nat Biotechnol. 2004. PMID: 15247925
-
Comparative genomic and transcriptomic characterization of the toxigenic marine dinoflagellate Alexandrium ostenfeldii.PLoS One. 2011;6(12):e28012. doi: 10.1371/journal.pone.0028012. Epub 2011 Dec 2. PLoS One. 2011. PMID: 22164224 Free PMC article.
-
An overview of transcription in dinoflagellates.Gene. 2022 Jun 30;829:146505. doi: 10.1016/j.gene.2022.146505. Epub 2022 Apr 18. Gene. 2022. PMID: 35447242 Review.
-
Biosynthesis and molecular genetics of polyketides in marine dinoflagellates.Mar Drugs. 2010 Mar 31;8(4):1011-48. doi: 10.3390/md8041011. Mar Drugs. 2010. PMID: 20479965 Free PMC article. Review.
Cited by
-
Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle.Sci Rep. 2016 Dec 22;6:39734. doi: 10.1038/srep39734. Sci Rep. 2016. PMID: 28004835 Free PMC article.
-
The Mechanism of Diarrhetic Shellfish Poisoning Toxin Production in Prorocentrum spp.: Physiological and Molecular Perspectives.Toxins (Basel). 2016 Sep 22;8(10):272. doi: 10.3390/toxins8100272. Toxins (Basel). 2016. PMID: 27669302 Free PMC article. Review.
-
Analysis of EST data of the marine protist Oxyrrhis marina, an emerging model for alveolate biology and evolution.BMC Genomics. 2014 Feb 11;15:122. doi: 10.1186/1471-2164-15-122. BMC Genomics. 2014. PMID: 24512041 Free PMC article.
-
Full-length transcriptome analysis of the bloom-forming dinoflagellate Akashiwo sanguinea by single-molecule real-time sequencing.Front Microbiol. 2022 Oct 17;13:993914. doi: 10.3389/fmicb.2022.993914. eCollection 2022. Front Microbiol. 2022. PMID: 36325025 Free PMC article.
-
An overview on the marine neurotoxin, saxitoxin: genetics, molecular targets, methods of detection and ecological functions.Mar Drugs. 2013 Mar 27;11(4):991-1018. doi: 10.3390/md11040991. Mar Drugs. 2013. PMID: 23535394 Free PMC article. Review.
References
-
- Hackett JD, Anderson DM, Erdner DL, Bhattacharya D. Dinoflagellates: a remarkable evolutionary experiment. Amer J Bot. 2004;91:1523–1534. - PubMed
-
- Rizzo PJ. The enigma of the dinoflagellate chromosome. J Protozool. 1991;38:246–252.
-
- LaJeunesse TC, Lambert G, Andersen RA, Coffroth MA, Galbraith DW. Symbiodinium (Pyrrhophyta) genome sizes (DNA content) are smallest among dinoflagellates. J Phycol. 2005;41:880–886. doi: 10.1111/j.0022-3646.2005.04231.x. - DOI
-
- Spector DL. Dinoflagellate Nuclei. In: Spector DL, editor. Dinoflagellates. New York: Academic Press, Inc; 1984. pp. 107–147.
-
- Beam J, Himes M. Dinoflagellate genetics. In: Spector D, editor. Dinoflagellates. New York: Academic Press, Inc; 1984. pp. 263–298.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

