Sequence and haplotype analysis supports HLA-C as the psoriasis susceptibility 1 gene
- PMID: 16642438
- PMCID: PMC1474031
- DOI: 10.1086/503821
Sequence and haplotype analysis supports HLA-C as the psoriasis susceptibility 1 gene
Abstract
Previous studies have narrowed the interval containing PSORS1, the psoriasis-susceptibility locus in the major histocompatibility complex (MHC), to an approximately 300-kb region containing HLA-C and at least 10 other genes. In an effort to identify the PSORS1 gene, we cloned and completely sequenced this region from both chromosomes of five individuals. Two of the sequenced haplotypes were associated with psoriasis (risk), and the other eight were clearly unassociated (nonrisk). Comparison of sequence of the two risk haplotypes identified a 298-kb region of homology, extending from just telomeric of HLA-B to the HCG22 gene, which was flanked by clearly nonhomologous regions. Similar haplotypes cloned from unrelated individuals had nearly identical sequence. Combinatorial analysis of exonic variations in the known genes of the candidate interval revealed that HCG27, PSORS1C3, OTF3, TCF19, HCR, STG, and HCG22 bore no alleles unique to risk haplotypes among the 10 sequenced haplotypes. SPR1 and SEEK1 both had messenger RNA alleles specific to risk haplotypes, but only HLA-C and CDSN yielded protein alleles unique to risk. The risk alleles of HLA-C and CDSN (HLA-Cw6 and CDSN*TTC) were genotyped in 678 families with early-onset psoriasis; 620 of these families were also typed for 34 microsatellite markers spanning the PSORS1 interval. Recombinant haplotypes retaining HLA-Cw6 but lacking CDSN*TTC were significantly associated with psoriasis, whereas recombinants retaining CDSN*TTC but lacking HLA-Cw6 were not associated, despite good statistical power. By grouping recombinants with similar breakpoints, the most telomeric quarter of the 298-kb candidate interval could be excluded with high confidence. These results strongly suggest that HLA-Cw6 is the PSORS1 risk allele that confers susceptibility to early-onset psoriasis.
Figures
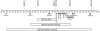

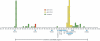


References
Web Resources
-
- Anthony Nolan Trust, http://www.anthonynolan.org.uk/HIG/ (for the IMGT/HLA sequence database)
-
- BLAST 2 Sequences, http://www.ncbi.nlm.nih.gov/blast/bl2seq/wblast2.cgi
-
- GenBank, http://www.ncbi.nih.gov/Genbank/ (see tables and for accession numbers)
References
-
- Gupta MA, Schork NJ, Gupta AK, Kirkby S, Ellis CN (1993) Suicidal ideation in psoriasis. Int J Dermatol 32:188–190 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

