On the accuracy of homology modeling and sequence alignment methods applied to membrane proteins
- PMID: 16648166
- PMCID: PMC1483079
- DOI: 10.1529/biophysj.106.082313
On the accuracy of homology modeling and sequence alignment methods applied to membrane proteins
Abstract
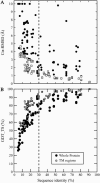
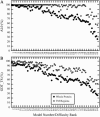
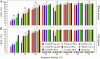
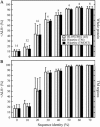
In this study, we investigate the extent to which techniques for homology modeling that were developed for water-soluble proteins are appropriate for membrane proteins as well. To this end we present an assessment of current strategies for homology modeling of membrane proteins and introduce a benchmark data set of homologous membrane protein structures, called HOMEP. First, we use HOMEP to reveal the relationship between sequence identity and structural similarity in membrane proteins. This analysis indicates that homology modeling is at least as applicable to membrane proteins as it is to water-soluble proteins and that acceptable models (with C alpha-RMSD values to the native of 2 A or less in the transmembrane regions) may be obtained for template sequence identities of 30% or higher if an accurate alignment of the sequences is used. Second, we show that secondary-structure prediction algorithms that were developed for water-soluble proteins perform approximately as well for membrane proteins. Third, we provide a comparison of a set of commonly used sequence alignment algorithms as applied to membrane proteins. We find that high-accuracy alignments of membrane protein sequences can be obtained using state-of-the-art profile-to-profile methods that were developed for water-soluble proteins. Improvements are observed when weights derived from the secondary structure of the query and the template are used in the scoring of the alignment, a result which relies on the accuracy of the secondary-structure prediction of the query sequence. The most accurate alignments were obtained using template profiles constructed with the aid of structural alignments. In contrast, a simple sequence-to-sequence alignment algorithm, using a membrane protein-specific substitution matrix, shows no improvement in alignment accuracy. We suggest that profile-to-profile alignment methods should be adopted to maximize the accuracy of homology models of membrane proteins.
Figures
Similar articles
-
Consensus alignment for reliable framework prediction in homology modeling.Bioinformatics. 2003 Sep 1;19(13):1682-91. doi: 10.1093/bioinformatics/btg211. Bioinformatics. 2003. PMID: 12967965
-
Structure-dependent sequence alignment for remotely related proteins.Bioinformatics. 2002 Dec;18(12):1658-65. doi: 10.1093/bioinformatics/18.12.1658. Bioinformatics. 2002. PMID: 12490451
-
Structure alignment of membrane proteins: Accuracy of available tools and a consensus strategy.Proteins. 2015 Sep;83(9):1720-32. doi: 10.1002/prot.24857. Epub 2015 Aug 1. Proteins. 2015. PMID: 26178143 Free PMC article.
-
Substitution scoring matrices for proteins - An overview.Protein Sci. 2020 Nov;29(11):2150-2163. doi: 10.1002/pro.3954. Epub 2020 Oct 12. Protein Sci. 2020. PMID: 32954566 Free PMC article. Review.
-
Homology Modeling of Transporter Proteins.Methods Mol Biol. 2023;2627:247-264. doi: 10.1007/978-1-0716-2974-1_14. Methods Mol Biol. 2023. PMID: 36959452 Review.
Cited by
-
Visualization of protein sequence space with force-directed graphs, and their application to the choice of target-template pairs for homology modelling.J Mol Graph Model. 2019 Nov;92:180-191. doi: 10.1016/j.jmgm.2019.07.014. Epub 2019 Jul 26. J Mol Graph Model. 2019. PMID: 31377535 Free PMC article.
-
Comparison, Analysis, and Molecular Dynamics Simulations of Structures of a Viral Protein Modeled Using Various Computational Tools.Bioengineering (Basel). 2023 Aug 24;10(9):1004. doi: 10.3390/bioengineering10091004. Bioengineering (Basel). 2023. PMID: 37760106 Free PMC article.
-
Computational studies of the binding site of alpha1A-adrenoceptor antagonists.J Mol Model. 2008 Oct;14(10):957-66. doi: 10.1007/s00894-008-0342-8. Epub 2008 Jul 15. J Mol Model. 2008. PMID: 18626669
-
The bacterial dicarboxylate transporter VcINDY uses a two-domain elevator-type mechanism.Nat Struct Mol Biol. 2016 Mar;23(3):256-63. doi: 10.1038/nsmb.3166. Epub 2016 Feb 1. Nat Struct Mol Biol. 2016. PMID: 26828963 Free PMC article.
-
Construction and validation of a homology model of the human voltage-gated proton channel hHV1.J Gen Physiol. 2013 Apr;141(4):445-65. doi: 10.1085/jgp.201210856. J Gen Physiol. 2013. PMID: 23530137 Free PMC article.
References
-
- Jones, D. T. 1998. Do transmembrane protein superfolds exist? FEBS Lett. 423:281–285. - PubMed
-
- Krogh, A., B. Larsson, G. von Heijne, and E. L. L. Sonnhammer. 2001. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305:567–580. - PubMed
-
- Drews, J. 2000. Drug discovery: a historical perspective. Science. 287:1960–1964. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

