Ion-mediated nucleic acid helix-helix interactions
- PMID: 16648172
- PMCID: PMC1483110
- DOI: 10.1529/biophysj.106.084285
Ion-mediated nucleic acid helix-helix interactions
Abstract
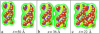
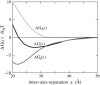
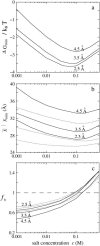
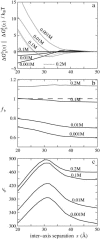
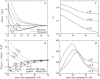
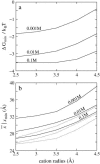
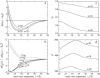
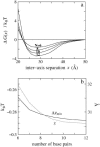
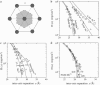
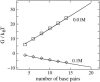
Salt ions are essential for the folding of nucleic acids. We use the tightly bound ion (TBI) model, which can account for the correlations and fluctuations for the ions bound to the nucleic acids, to investigate the electrostatic free-energy landscape for two parallel nucleic acid helices in the solution of added salt. The theory is based on realistic atomic structures of the helices. In monovalent salt, the helices are predicted to repel each other. For divalent salt, while the mean-field Poisson-Boltzmann theory predicts only the repulsion, the TBI theory predicts an effective attraction between the helices. The helices are predicted to be stabilized at an interhelix distance approximately 26-36 A, and the strength of the attractive force can reach -0.37 k(B)T/bp for helix length in the range of 9-12 bp. Both the stable helix-helix distance and the strength of the attraction are strongly dependent on the salt concentration and ion size. With the increase of the salt concentration, the helix-helix attraction becomes stronger and the most stable helix-helix separation distance becomes smaller. For divalent ions, at very high ion concentration, further addition of ions leads to the weakening of the attraction. Smaller ion size causes stronger helix-helix attraction and stabilizes the helices at a shorter distance. In addition, the TBI model shows that a decrease in the solvent dielectric constant would enhance the ion-mediated attraction. The theoretical findings from the TBI theory agree with the experimental measurements on the osmotic pressure of DNA array as well as the results from the computer simulations.
Figures
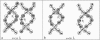

References
-
- Bloomfield, V. A., D. M. Crothers, and I. Tinoco, Jr. 2000. Nucleic Acids: Structure, Properties and Functions. University Science Books, Sausalito, CA.
-
- Tinoco, I., and C. Bustamante. 1999. How RNA folds. J. Mol. Biol. 293:271–281. - PubMed
-
- Anderson, C. F., and M. T. Record, Jr. 1995. Salt-nucleic acid interactions. Annu. Rev. Phys. Chem. 46:657–700. - PubMed
-
- Sosnick, T. R., and T. Pan. 2003. RNA folding: models and perspectives. Curr. Opin. Struct. Biol. 13:309–316. - PubMed
-
- Woodson, S. A. 2005. Metal ions and RNA folding: a highly charged topic with a dynamic future. Curr. Opin. Chem. Biol. 9:104–109. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

