SirT2 is a histone deacetylase with preference for histone H4 Lys 16 during mitosis
- PMID: 16648462
- PMCID: PMC1472900
- DOI: 10.1101/gad.1412706
SirT2 is a histone deacetylase with preference for histone H4 Lys 16 during mitosis
Abstract
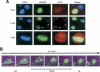
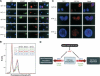
The mammalian cytoplasmic protein SirT2 is a member of the Sir2 family of NAD+-dependent protein deacetylases involved in caloric restriction-dependent life span extension. We found that SirT2 and its yeast counterpart Hst2 have a strong preference for histone H4K16Ac in their deacetylation activity in vitro and in vivo. We have pinpointed the decrease in global levels of H4K16Ac during the mammalian cell cycle to the G2/M transition that coincides with SirT2 localization on chromatin. Mouse embryonic fibroblasts (MEFs) deficient for SirT2 show higher levels of H4K16Ac in mitosis, in contrast to the normal levels exhibited by SirT1-deficient MEFs. The enzymatic conversion of H4K16Ac to its deacetylated form may be pivotal to the formation of condensed chromatin. Thus, SirT2 is a major contributor to this enzymatic conversion at the time in the cell's life cycle when condensed chromatin must be generated anew.
Figures


Similar articles
-
The molecular biology of mammalian SIRT proteins: SIRT2 in cell cycle regulation.Cell Cycle. 2007 May 2;6(9):1011-8. doi: 10.4161/cc.6.9.4219. Epub 2007 May 30. Cell Cycle. 2007. PMID: 17457050 Review.
-
Substrate specificity and kinetic mechanism of the Sir2 family of NAD+-dependent histone/protein deacetylases.Biochemistry. 2004 Aug 3;43(30):9877-87. doi: 10.1021/bi049592e. Biochemistry. 2004. PMID: 15274642
-
Histone deacetylase Rpd3 antagonizes Sir2-dependent silent chromatin propagation.Nucleic Acids Res. 2009 Jun;37(11):3699-713. doi: 10.1093/nar/gkp233. Epub 2009 Apr 16. Nucleic Acids Res. 2009. PMID: 19372273 Free PMC article.
-
The human Sir2 ortholog, SIRT2, is an NAD+-dependent tubulin deacetylase.Mol Cell. 2003 Feb;11(2):437-44. doi: 10.1016/s1097-2765(03)00038-8. Mol Cell. 2003. PMID: 12620231
-
NAD+-dependent deacetylation of H4 lysine 16 by class III HDACs.Oncogene. 2007 Aug 13;26(37):5505-20. doi: 10.1038/sj.onc.1210617. Oncogene. 2007. PMID: 17694090 Review.
Cited by
-
Lysine acetylation of cytoskeletal proteins: Emergence of an actin code.J Cell Biol. 2020 Dec 7;219(12):e202006151. doi: 10.1083/jcb.202006151. J Cell Biol. 2020. PMID: 33044556 Free PMC article. Review.
-
Exploring the Multi-Faceted Role of Sirtuins in Glioblastoma Pathogenesis and Targeting Options.Int J Mol Sci. 2022 Oct 25;23(21):12889. doi: 10.3390/ijms232112889. Int J Mol Sci. 2022. PMID: 36361680 Free PMC article. Review.
-
Quantitative acetylome analysis reveals the roles of SIRT1 in regulating diverse substrates and cellular pathways.Mol Cell Proteomics. 2012 Oct;11(10):1048-62. doi: 10.1074/mcp.M112.019547. Epub 2012 Jul 23. Mol Cell Proteomics. 2012. PMID: 22826441 Free PMC article.
-
SIRT2 is a tumor suppressor that connects aging, acetylome, cell cycle signaling, and carcinogenesis.Transl Cancer Res. 2012 Jun 1;1(1):15-21. Epub 2012 May 22. Transl Cancer Res. 2012. PMID: 22943040 Free PMC article.
-
SIRT2 inhibition achieves neuroprotection by decreasing sterol biosynthesis.Proc Natl Acad Sci U S A. 2010 Apr 27;107(17):7927-32. doi: 10.1073/pnas.1002924107. Epub 2010 Apr 8. Proc Natl Acad Sci U S A. 2010. PMID: 20378838 Free PMC article.
References
-
- Bae N.S., Swanson M.J., Vassilev A., Howard B.H. Human histone deacetylase SIRT2 interacts with the homeobox transcription factor HOXA10. J. Biochem. (Tokyo) 2004;135:695–700. - PubMed
-
- Belyaev N.D., Houben A., Baranczewski P., Schubert I. Histone H4 acetylation in plant heterochromatin is altered during the cell cycle. Chromosoma. 1997;106:193–197. - PubMed
-
- Braunstein M., Rose A.B., Holmes S.G., Allis C.D., Broach J.R. Transcriptional silencing in yeast is associated with reduced nucleosome acetylation. Genes & Dev. 1993;7:592–604. - PubMed
-
- Chuikov S., Kurash J.K., Wilson J.R., Xiao B., Justin N., Ivanov G.S., McKinney K., Tempst P., Prives C., Gamblin S.J., et al. Regulation of p53 activity through lysine methylation. Nature. 2004;432:353–360. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
