Transcription regulatory elements are punctuation marks for DNA replication
- PMID: 16670199
- PMCID: PMC1464333
- DOI: 10.1073/pnas.0601127103
Transcription regulatory elements are punctuation marks for DNA replication
Abstract
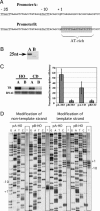
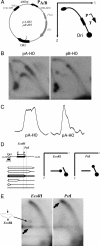
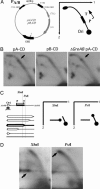
Collisions between DNA replication and transcription significantly affect genome organization, regulation, and stability. Previous studies have described collisions between replication forks and elongating RNA polymerases. Although replication collisions with the transcription-initiation or -termination complexes are potentially even more important because most genes are not actively transcribed during DNA replication, their existence and mechanisms remained unproven. To address this matter, we have designed a bacterial promoter that binds RNA polymerase and maintains it in the initiating mode by precluding the transition into the elongation mode. By using electrophoretic analysis of replication intermediates, we have found that this steadfast transcription-initiation complex inhibits replication fork progression in an orientation-dependent manner during head-on collisions. Transcription terminators also appeared to attenuate DNA replication, but in the opposite, codirectional orientation. Thus, transcription regulatory signals may serve as "punctuation marks" for DNA replication in vivo.
Conflict of interest statement
Conflict of interest statement: No conflicts declared.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

