Conversion of trypsin to a functional threonine protease
- PMID: 16672242
- PMCID: PMC2242550
- DOI: 10.1110/ps.062179006
Conversion of trypsin to a functional threonine protease
Abstract
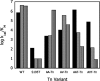
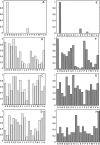
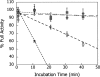
The hydroxyl group of a serine residue at position 195 acts as a nucleophile in the catalytic mechanism of the serine proteases. However, the chemically similar residue, threonine, is rarely used in similar functional context. Our structural modeling suggests that the Ser 195 --> Thr trypsin variant is inactive due to negative steric interaction between the methyl group on the beta-carbon of Thr 195 and the disulfide bridge formed by cysteines 42 and 58. By simultaneously truncating residues 42 and 58 and substituting Ser 195 with threonine, we have successfully converted the classic serine protease trypsin to a functional threonine protease. Substitution of residue 42 with alanine and residue 58 with alanine or valine in the presence of threonine 195 results in trypsin variants that are 10(2) -10(4) -fold less active than wild type in kcat/KM but >10(6)-fold more active than the Ser 195 --> Thr single variant. The substitutions do not alter the substrate specificity of the enzyme in the P1'- P4' positions. Removal of the disulfide bridge decreases the overall thermostability of the enzyme, but it is partially rescued by the presence of threonine at position 195.
Figures

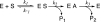
Similar articles
-
Why Ser and not Thr brokers catalysis in the trypsin fold.Biochemistry. 2015 Feb 24;54(7):1457-64. doi: 10.1021/acs.biochem.5b00014. Epub 2015 Feb 11. Biochemistry. 2015. PMID: 25664608 Free PMC article.
-
The role of the Cys191-Cys220 disulfide bond in trypsin: new targets for engineering substrate specificity.Protein Eng. 1997 Apr;10(4):405-11. doi: 10.1093/protein/10.4.405. Protein Eng. 1997. PMID: 9194165
-
Engineering the S1' subsite of trypsin: design of a protease which cleaves between dibasic residues.Biochemistry. 1998 Aug 18;37(33):11434-40. doi: 10.1021/bi980842z. Biochemistry. 1998. PMID: 9708978
-
Structure-function relationship of serine protease-protein inhibitor interaction.Acta Biochim Pol. 2001;48(2):419-28. Acta Biochim Pol. 2001. PMID: 11732612 Review.
-
Trypsin: a case study in the structural determinants of enzyme specificity.Biol Chem. 1996 Jul-Aug;377(7-8):465-70. Biol Chem. 1996. PMID: 8922280 Review.
Cited by
-
The active site region plays a critical role in Na+ binding to thrombin.J Biol Chem. 2022 Jan;298(1):101458. doi: 10.1016/j.jbc.2021.101458. Epub 2021 Nov 30. J Biol Chem. 2022. PMID: 34861239 Free PMC article.
-
TMEM164 is an acyltransferase that forms ferroptotic C20:4 ether phospholipids.Nat Chem Biol. 2023 Mar;19(3):378-388. doi: 10.1038/s41589-022-01253-7. Epub 2023 Feb 13. Nat Chem Biol. 2023. PMID: 36782012 Free PMC article.
-
NBCZone: Universal three-dimensional construction of eleven amino acids near the catalytic nucleophile and base in the superfamily of (chymo)trypsin-like serine fold proteases.Int J Biol Macromol. 2020 Jun 15;153:399-411. doi: 10.1016/j.ijbiomac.2020.03.025. Epub 2020 Mar 6. Int J Biol Macromol. 2020. PMID: 32151723 Free PMC article.
-
Engineering trypsin for inhibitor resistance.Protein Sci. 2015 Sep;24(9):1463-74. doi: 10.1002/pro.2732. Epub 2015 Jul 7. Protein Sci. 2015. PMID: 26106067 Free PMC article.
-
Codon switching of conserved Ser residues in coagulation and fibrinolytic proteases.J Thromb Haemost. 2024 Sep;22(9):2495-2501. doi: 10.1016/j.jtha.2024.05.021. Epub 2024 May 29. J Thromb Haemost. 2024. PMID: 38821294
References
-
- Baumeister W. and Lupas A. 1997. The proteasome Curr. Opin. Struct. Biol. 7 273–278. - PubMed
-
- Betzel C., Gourinath S., Kumar P., Kaur P., Perbandt M., Eschenburg S., Singh T.P. 2001. Structure of a serine protease proteinase K from Tritirachium album limber at 0.98 Å resolution Biochemistry 40 3080–3088. - PubMed
-
- Bode W., Turk D., Karshikov A. 1992. The refined 1.9-Å X-ray crystal structure of D-Phe-Pro-Arg chloromethylketone-inhibited human α-thrombin: Structure analysis, overall structure, electrostatic properties, detailed active-site geometry, and structure–function relationships Protein Sci. 1 426–471. - PMC - PubMed
-
- Brannigan J.A., Dodson G., Duggleby H.J., Moody P.C., Smith J.L., Tomchick D.R., Murzin A.G. 1995. A protein catalytic framework with an N-terminal nucleophile is capable of self-activation Nature 378 416–419. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

