The structural basis of paramyxovirus invasion
- PMID: 16678421
- PMCID: PMC7119026
- DOI: 10.1016/j.tim.2006.04.004
The structural basis of paramyxovirus invasion
Abstract
To deliver their genetic material into host cells, enveloped viruses have surface glycoproteins that actively cause the fusion of the viral and cellular membranes. Recently determined X-ray crystal structures of the paramyxovirus fusion (F) protein in its pre-fusion and post-fusion conformations reveal the dramatic structural transformation that this protein undergoes while causing membrane fusion. Conformational changes in key regions of the F protein suggest the mechanism by which the F protein is activated and refolds.
Figures
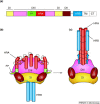
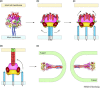
References
-
- Crennell S. Crystal structure of the multifunctional paramyxovirus hemagglutinin-neuraminidase. Nat. Struct. Biol. 2000;7:1068–1074. - PubMed
-
- Lawrence M.C. Structure of the haemagglutinin-neuraminidase from human parainfluenza virus type III. J. Mol. Biol. 2004;335:1343–1357. - PubMed
-
- Yuan P. Structural studies of the parainfluenza virus 5 hemagglutinin-neuraminidase tetramer in complex with its receptor, sialyllactose. Structure. 2005;13:803–815. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

