Every enhancer works with every promoter for all the combinations tested: could new regulatory pathways evolve by enhancer shuffling?
- PMID: 1668145
- PMCID: PMC5952201
Every enhancer works with every promoter for all the combinations tested: could new regulatory pathways evolve by enhancer shuffling?
Abstract
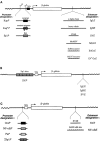
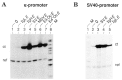
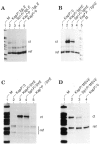
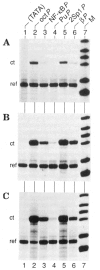
The promoters and enhancers of cell type-specific genes are often conserved in evolution, and hence one might expect that a given enhancer has evolved to work best with its own promoter. While this expectation may be realized in some cases, we have not found evidence for it. A total of 27 combinations of different promoters and enhancers were tested by transfection into cultured cells. We found that the relative efficiency of the enhancers is approximately the same, irrespective of the type of promoter used, i.e., there was no strong preference for any given enhancer/promoter combination. Notably, we do not see particularly strong transcription when the immunoglobulin kappa enhancer (or the immunoglobulin heavy chain enhancer) is used to activate a kappa gene promoter. We propose that a generally permissive enhancer/promoter interaction is of evolutionary benefit for higher eukaryotes: by enhancer shuffling, genes could be easily brought under a new type of inducibility/cell type specificity.
Figures
Similar articles
-
Synergism between immunoglobulin enhancers and promoters.Nature. 1986 Jul 24-30;322(6077):383-5. doi: 10.1038/322383a0. Nature. 1986. PMID: 3016550
-
Kappa immunoglobulin promoters and enhancers display developmentally controlled interactions.Nucleic Acids Res. 1993 Oct 25;21(21):4941-7. doi: 10.1093/nar/21.21.4941. Nucleic Acids Res. 1993. PMID: 8177743 Free PMC article.
-
Functional similarities between human immunodeficiency virus type 1 and simian virus 40 kappa B proto-enhancers.J Virol. 1991 May;65(5):2200-10. doi: 10.1128/JVI.65.5.2200-2210.1991. J Virol. 1991. PMID: 1850006 Free PMC article.
-
Cell-type specific regulation of a kappa immunoglobulin gene by promoter and enhancer elements.Immunol Rev. 1986 Feb;89:49-68. doi: 10.1111/j.1600-065x.1986.tb01472.x. Immunol Rev. 1986. PMID: 3081432 Review. No abstract available.
-
Long distance relationships: enhancer-promoter communication and dynamic gene transcription.Biochim Biophys Acta. 2012 Nov-Dec;1819(11-12):1217-27. doi: 10.1016/j.bbagrm.2012.10.008. Epub 2012 Nov 1. Biochim Biophys Acta. 2012. PMID: 23124110 Review.
Cited by
-
Spermiogenesis and Male Fertility Require the Function of Suppressor of Hairy-Wing in Somatic Cyst Cells of Drosophila.Genetics. 2018 Jul;209(3):757-772. doi: 10.1534/genetics.118.301088. Epub 2018 May 8. Genetics. 2018. PMID: 29739818 Free PMC article.
-
Lack of the Drosophila BEAF insulator proteins alters regulation of genes in the Antennapedia complex.Mol Genet Genomics. 2011 Feb;285(2):113-23. doi: 10.1007/s00438-010-0591-y. Epub 2010 Dec 4. Mol Genet Genomics. 2011. PMID: 21132442
-
A 61 bp enhancer element of the tobacco beta-1,3-glucanase B gene interacts with one or more regulated nuclear proteins.Plant Mol Biol. 1993 Jan;21(1):121-31. doi: 10.1007/BF00039623. Plant Mol Biol. 1993. PMID: 8425042
-
Adaptive variation in lactate dehydrogenase-B gene expression: role of a stress-responsive regulatory element.Proc Natl Acad Sci U S A. 2000 Jun 6;97(12):6597-602. doi: 10.1073/pnas.97.12.6597. Proc Natl Acad Sci U S A. 2000. PMID: 10841559 Free PMC article.
-
An initiator element mediates autologous downregulation of the human type A gamma -aminobutyric acid receptor beta 1 subunit gene.Proc Natl Acad Sci U S A. 2000 Jul 18;97(15):8600-5. doi: 10.1073/pnas.97.15.8600. Proc Natl Acad Sci U S A. 2000. PMID: 10900018 Free PMC article.
References
-
- Abel T. and Maniatis T. (1990), Nature 341, 24–25. - PubMed
-
- Atchison M. L. and Perry R. P. (1987), Cell 48, 121–128. - PubMed
-
- Banerji J., Rusconi S., and Schaffner W. (1981), Cell 27, 299–308. - PubMed
-
- Banerji J., Olson L., and Schaffner W. (1983), Cell 33, 729–740. - PubMed
-
- Benoist C. and Chambon P. (1981), Nature 290, 310–315. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
