Early events in protein folding explored by rapid mixing methods
- PMID: 16683757
- PMCID: PMC2556641
- DOI: 10.1021/cr040430y
Early events in protein folding explored by rapid mixing methods
Figures


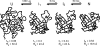
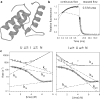
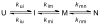
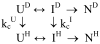
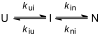
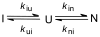
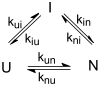
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

