Cathepsin L stabilizes the histone modification landscape on the Y chromosome and pericentromeric heterochromatin
- PMID: 16705169
- PMCID: PMC1489105
- DOI: 10.1128/MCB.00135-06
Cathepsin L stabilizes the histone modification landscape on the Y chromosome and pericentromeric heterochromatin
Abstract
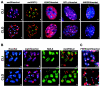
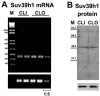
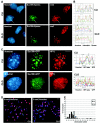
Posttranslational histone modifications and histone variants form a unique epigenetic landscape on mammalian chromosomes where the principal epigenetic heterochromatin markers, trimethylated histone H3(K9) and the histone H2A.Z, are inversely localized in relation to each other. Trimethylated H3(K9) marks pericentromeric constitutive heterochromatin and the male Y chromosome, while H2A.Z is dramatically reduced at these chromosomal locations. Inactivation of a lysosomal and nuclear protease, cathepsin L, causes a global redistribution of epigenetic markers. In cathepsin L knockout cells, the levels of trimethylated H3(K9) decrease dramatically, concomitant with its relocation away from heterochromatin, and H2A.Z becomes enriched at pericentromeric heterochromatin and the Y chromosome. This change is also associated with global relocation of heterochromatin protein HP1 and histone H3 methyltransferase Suv39h1 away from constitutive heterochromatin; however, it does not affect DNA methylation or chromosome segregation, phenotypes commonly associated with impaired histone H3(K9) methylation. Therefore, the key constitutive heterochromatin determinants can dynamically redistribute depending on physiological context but still maintain the essential function(s) of chromosomes. Thus, our data show that cathepsin L stabilizes epigenetic heterochromatin markers on pericentromeric heterochromatin and the Y chromosome through a novel mechanism that does not involve DNA methylation or affect heterochromatin structure and operates on both somatic and sex chromosomes.
Figures


References
-
- Al-Khunaizi, M., C. J. Luke, Y. S. Askew, S. C. Pak, D. J. Askew, S. Cataltepe, D. Miller, D. R. Mills, C. Tsu, D. Bromme, J. A. Irving, J. C. Whisstock, and G. A. Silverman. 2002. The serpin SQN-5 is a dual mechanistic-class inhibitor of serine and cysteine proteinases. Biochemistry 41:3189-3199. - PubMed
-
- Bannister, A. J., P. Zegerman, J. F. Partridge, E. A. Miska, J. O. Thomas, R. C. Allshire, and T. Kouzarides. 2001. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410:120-124. - PubMed
-
- Barrett, A. J., and H. Kirschke. 1981. Cathepsin B, cathepsin H, and cathepsin L. Methods Enzymol. 80(Pt C):535-561. - PubMed
-
- Barton, S. C., K. L. Arney, W. Shi, A. Niveleau, R. Fundele, M. A. Surani, and T. Haaf. 2001. Genome-wide methylation patterns in normal and uniparental early mouse embryos. Hum. Mol. Genet. 10:2983-2987. - PubMed
-
- Bean, C. J., P. A. Hunt, E. A. Millie, and T. J. Hassold. 2001. Analysis of a malsegregating mouse Y chromosome: evidence that the earliest cleavage divisions of the mammalian embryo are non-disjunction-prone. Hum. Mol. Genet. 10:963-972. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
